99 KiB
Experimental Validation
Roberto Olayo Alarcon and Medina Bajramovic 27/03/2024
In this file, we look at the output from the experimental validation.
library(tidyverse)
library(readxl)
Prepare directories.
RAW_DATA_DIR <- "../raw_data/"
OUTPUT_DIR <- "../data/06.experimental_validation"
# Create output dir
if(!dir.exists(OUTPUT_DIR)){
dir.create(OUTPUT_DIR)
}
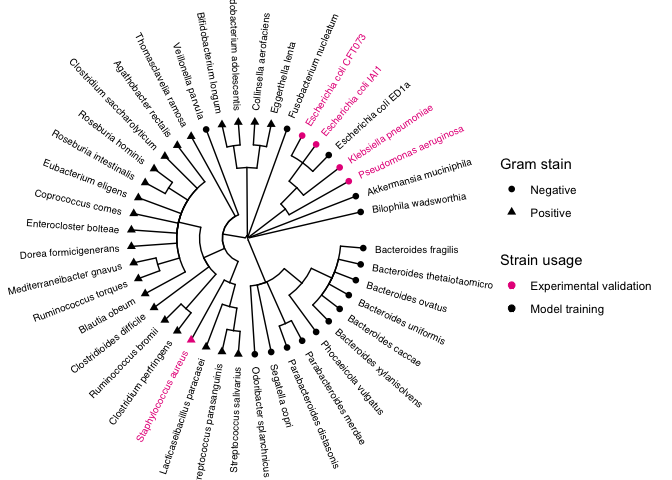
Taxonimic tree of species
library(ggtree)
First we can read the taxanomic tree data.
taxtree <- treeio::read.newick(file.path(RAW_DATA_DIR, "maier_microbiome/phyliptree_wPath.phy"))
Read strain metadata.
strain_metadata <- read_excel(file.path(RAW_DATA_DIR, "maier_microbiome/strain_info_SF2.xlsx"),
skip = 2)
# Gather Gram strains
strain_gram <- strain_metadata %>% select(Species, `Gram stain`) %>%
filter(!is.na(Species))
Now we can clean up the strain names from NCBI.
process_strain_name <- function(strain){
# Pre-process strain name
strain_red <- str_remove_all(strain, "'")
strain_red <- str_remove_all(strain_red, "\\[")
strain_red <- str_remove_all(strain_red, "\\]")
#print(strain_red)
if(str_starts(strain_red, "Escherichia")){
return(strain_red)
}else{
strain_red <- paste(str_split(strain_red, " ", simplify = TRUE)[, 1:2], collapse = " ")
return(strain_red)
}
}
taxtree$tip.label <- sapply(taxtree$tip.label, process_strain_name)
Now we can assign some labels to these taxons.
validation_strains <- c("Escherichia coli CFT073",
"Pseudomonas aeruginosa",
"Staphylococcus aureus",
"Klebsiella pneumoniae",
"Escherichia coli IAI1")
tip.use <- data.frame(Species=taxtree$tip.label) %>%
mutate(strain_use = if_else(Species %in% validation_strains, "Experimental validation", "Model training")) %>%
rename("label" = "Species")
# Add gram stain information
tip.gram <- data.frame(Species=taxtree$tip.label) %>%
left_join(strain_gram, by="Species") %>%
# Manual addition of some gram stain
mutate(`Gram stain` = case_when(Species %in% c("Escherichia coli CFT073",
"Pseudomonas aeruginosa",
"Klebsiella pneumoniae",
"Escherichia coli IAI1",
"Fusobacterium nucleatum",
"Segatella copri",
"Phocaeicola vulgatus") ~ "negative",
Species %in% c("Staphylococcus aureus",
"Clostridioides difficile",
"Bifidobacterium longum",
"Thomasclavelia ramosa",
"Agathobacter rectalis",
"Enterocloster bolteae",
"Mediterraneibacter gnavus",
"Lacticaseibacillus paracasei") ~ "positive",
TRUE ~ `Gram stain`),
`Gram stain` = str_to_title(`Gram stain`)) %>%
rename("label" = "Species")
Now we can plot the tree
p.tree <- ggtree(taxtree, branch.length = "none", layout = "circular", ladderize = FALSE) %<+% tip.use %<+% tip.gram +
geom_tippoint(aes(color=strain_use, shape=`Gram stain`), size=2) +
geom_tiplab(size=2.6, offset = .5, aes(angle=angle, color=strain_use))+
scale_color_manual(values=c(`Model training` = "black", `Experimental validation`= "#e7298a")) +
scale_shape_manual(values=c(Negative=16, Positive=17)) +
hexpand(ratio=1) +
vexpand(ratio = 1.5) +
xlim(-.1, 8) +
labs(color="Strain usage")
p.tree
ggsave(plot=p.tree, filename = file.path(OUTPUT_DIR, "taxtree_circle.pdf"),
dpi=300, width = 19, height = 15, units = "cm")
ggsave(plot=p.tree, filename = file.path(OUTPUT_DIR, "taxtree_circle.png"),
dpi=300, width = 19, height = 15, units = "cm")
Read Optical Density.
Here we process the raw OD measurements from the machine. We perform baseline correction and ignore first timepoint.
read_od_files <- function(file_name, input_directory, baseline_t=c(2, 3, 4), clip_lower=TRUE){
#' Function to read and preprocess optical density (OD) files.
#'
#' @param file_name A character string specifying the name of the OD file.
#' @param input_directory A character string specifying the directory where the OD file is located.
#' @param baseline_t An optional numeric vector specifying the timepoints to be used for baseline correction. Defaults to c(2, 3, 4).
#' @param clip_lower An optional logical value indicating whether to clip negative values to zero. Defaults to TRUE.
#'
#' @return A data frame containing the preprocessed OD data in long format, with additional columns for timepoint, time spent in the incubator (in hours), well identifier, OD readings, file origin, organism, and replicate.
#'
#' The function reads an OD file from the specified directory, performs baseline correction, clips lower values (if specified),
#' adds time-related columns, transforms the data into long format, and adds additional information such as file origin, organism, and replicate.
#' The preprocessed data is returned as a data frame.
#'
# Read file
file_path <- file.path(input_directory, file_name)
timeseries_df <- read.table(file_path, sep='\t', header = TRUE) %>%
column_to_rownames("Time")
# Baseline correction. We look for the minimum of the timepoints specified for each well
well_minimums <- timeseries_df %>%
# Only look at specicied timepoints
slice(baseline_t) %>%
# Gather the minimum of each well
summarise(across(where(is.numeric), min)) %>%
unlist()
# Subtract baseline
timeseries_df <- sweep(timeseries_df, 2, well_minimums)
# Clip values
if(clip_lower){
timeseries_df[timeseries_df < 0] <- 0
}
# Add timepoint columns
timeseries_df <- timeseries_df %>%
# Add Time column
rownames_to_column("Time") %>%
# Add timepoint
mutate(timepoint = 1:n())
# Add time spent in incubator (in hours)
## Add time 0
t0 <- strptime(timeseries_df$Time[1], format = "%m/%d/%Y %H:%M:%S", tz = "CET")
timeseries_df <- timeseries_df %>%
## Format time-stamps
mutate(time_format = strptime(Time, format = "%m/%d/%Y %H:%M:%S", tz = "CET"),
# Time spent in incubator
time_spent_h = as.numeric(time_format - t0, units="hours")) %>%
select(-time_format)
# Long format
timeseries.long <- timeseries_df %>%
pivot_longer(cols = -c(timepoint, Time, time_spent_h), names_to = "well", values_to = "od")
# Add information
## Join ecoli strains
assay.id <- if_else(str_count(file_name, "_") == 3, str_remove(file_name, "_"), file_name)
assay.id <- str_remove(assay.id, "_OD.tsv.gz")
timeseries.long <- timeseries.long %>%
mutate(file_origin = file_name,
organism_replicate = assay.id) %>%
separate(organism_replicate, sep = "_", into = c("organism", "replicate"))
return(timeseries.long)
}
Now we iterate over all OD files
# Directory for od files
input_directory <- file.path(RAW_DATA_DIR, "experimental_validation", "od_files")
input_files <- list.files(input_directory)
# Process OD
complete_od <- bind_rows(lapply(input_files, read_od_files, input_directory=input_directory))
# Write processed OD
write_tsv(x=complete_od, file = file.path(OUTPUT_DIR, "complete_od.tsv.gz"))
Now we read the metadata
well_metadata <- read_excel(file.path(RAW_DATA_DIR, "experimental_validation", "lib_map.xlsx")) %>%
select(-well_code) %>%
mutate(concentration = as.numeric(concentration),
dmso_percentage = as.numeric(dmso_percentage))
What is important to note here is that now, each well refers to a specific Compound - Concentration. At the same time, we have an additional column that tells us the amount of DMSO used for the specific preparation.
This validation plate has 96 wells in total. We test this setup against 5 microbial strains in triplicate. This means that we will end up having a grand total of 96 x 5 x 3 = 1,440 individual growth curves. On top of this, Escherichia coli IAI was tested twice against this plate, meaning that we have an additional 288 curves. This gives us a total of 1,728 growth curves.
Now that we have all of the data, we can add the metadata to our growth data.
# Add information
complete_od <- complete_od %>%
select(-c(Time, file_origin)) %>%
left_join(well_metadata, by="well") %>%
# Remove noisy timepoints.
filter(timepoint >= 2) %>%
mutate(curve_id = paste(organism, well, replicate, sep="_"))
# Count number of curves
complete_od %>%
select(curve_id) %>%
n_distinct()
## [1] 1728
Explore experiments.
Now, we can get a general idea of the results from our experiments!
all_data_plot <- complete_od %>%
filter(compound != "DMSO") %>%
ggplot(aes(x=time_spent_h, y=od, group=curve_id, color=log2(concentration))) +
geom_line(linewidth=0.2) +
#scale_color_distiller(palette = "PuRd") +
facet_grid(organism~compound) +
theme_light() +
theme(legend.position = "bottom") +
ggtitle("Experimental results - Compound x Organism")
all_data_plot
From this we can see that Visomitin is active against Gram negative and Gram positive bacteria, as we predicted. This is a very neat result! Unfortunately, Visomitin is one name this compound has. It’s generic name is SKQ1, and this has already been shown to be active against a variety of bacterial species Nazarov P.A., 2017.
We also see that Staphylococcus aureus is suceptible to Elivitegravir and also sensitive to Opicapone. There is also some indications that Ebastine affects growth. Now we can take a closer look at each compound individually.
Baseline growth (Water and DMSO).
It’s worth to take a closer look at growth under DMSO and Water to look for outliers.
DMSO
complete_od %>%
filter(compound == "DMSO") %>%
mutate(Conc = factor(concentration, levels=sort(unique(concentration)))) %>%
ggplot(aes(x=timepoint, y=od, group=curve_id, color=concentration)) +
geom_line(linewidth=0.25) +
#scale_color_brewer(palette = "Paired") +
scale_color_distiller(palette = "PuRd") +
geom_vline(xintercept = 9, color="white", linetype="longdash")+
facet_wrap(~organism, nrow = 2, ncol = 4) +
theme_dark()
It seems like DMSO does have an effect on the growth of our Gram negative strains!
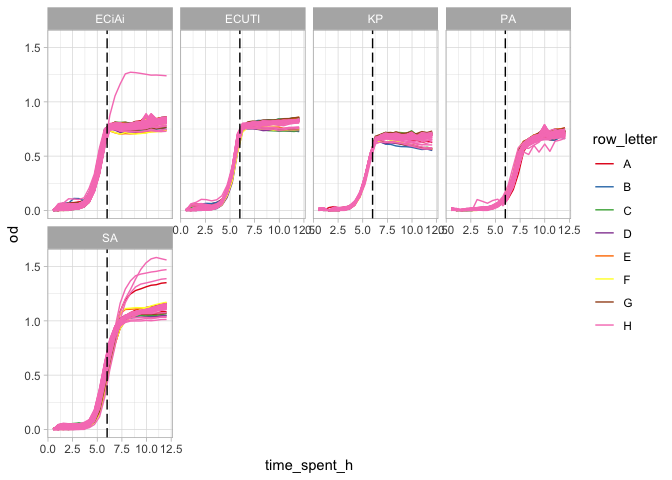
Water
complete_od %>%
filter(compound == "Water") %>%
mutate(row_letter = sapply(well, function(x){str_split(x, "", simplify = TRUE)[,1]})) %>%
ggplot(aes(x=time_spent_h, y=od, group=curve_id, color=row_letter)) +
geom_line() +
geom_vline(xintercept = 6, color="black", linetype="longdash") +
scale_color_brewer(palette = "Set1") +
facet_wrap(~organism, nrow = 2, ncol = 4) +
theme_light()
Seems like there might be some outliers with overgrowth for S. aureus, P. aeruginosa, and E. coli IAi.
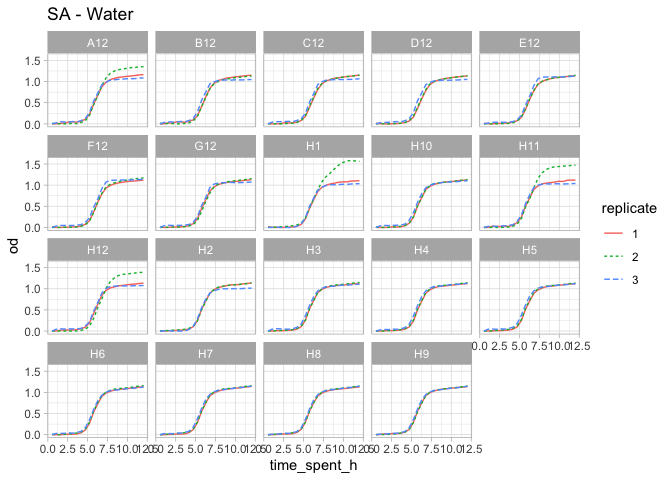
complete_od %>%
filter(compound == "Water", organism=="SA") %>%
mutate(row_letter = sapply(well, function(x){str_split(x, "", simplify = TRUE)[,1]})) %>%
#filter(row_letter == "H") %>%
ggplot(aes(x=time_spent_h, y=od, group=curve_id, color=replicate, linetype=replicate)) +
geom_line() +
facet_wrap(~well) +
theme_light() +
ggtitle("SA - Water")
complete_od %>%
filter(compound == "Water", organism=="PA") %>%
mutate(row_letter = sapply(well, function(x){str_split(x, "", simplify = TRUE)[,1]})) %>%
#filter(row_letter == "H") %>%
ggplot(aes(x=time_spent_h, y=od, group=curve_id, color=replicate, linetype=replicate)) +
geom_line() +
facet_wrap(~well) +
theme_light() +
ggtitle("PA - Water")

complete_od %>%
filter(compound == "Water", organism=="ECiAi") %>%
mutate(row_letter = sapply(well, function(x){str_split(x, "", simplify = TRUE)[,1]})) %>%
#filter(row_letter == "H") %>%
ggplot(aes(x=time_spent_h, y=od, group=curve_id, color=replicate, linetype=replicate)) +
geom_line() +
facet_wrap(~well) +
theme_light() +
ggtitle("ECiAi - Water")
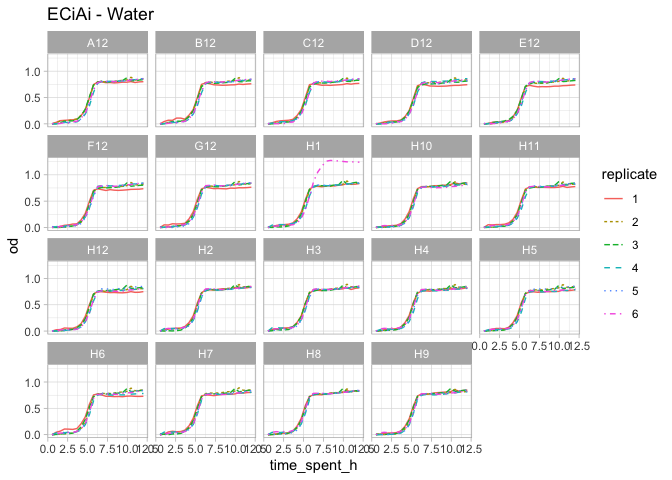
Given the plots above, I think it makes sense to remove the following curves and consider them as outliers.
outlier_water <- c("SA_H1_2", "SA_H11_2", "SA_H12_2", "SA_A12_2", "ECiAi_H1_6", "PA_H7_3")
complete_od <- complete_od %>%
filter(!(curve_id %in% outlier_water))
Visualize Concentration X Organism panel.
In this section we explore the effects of each compound.
# Create output panel directory
OUTPUT_PANEL_PLOTS_DIR <- file.path(OUTPUT_DIR, "organism_concentration_plots")
if(!dir.exists(OUTPUT_PANEL_PLOTS_DIR)){
dir.create(OUTPUT_PANEL_PLOTS_DIR)
}
# THIS function looks a bit more specifically at a given compound_concentration combination. Here we do give
# a bit more attention when selecting the correct DMSO curves and the correct Water curves.
gather_baseline <- function(cname, conc_i, growth_df, info_df){
#' Gather baseline data for a specific compound and concentration.
#'
#' This function retrieves baseline data necessary for calculating the mean growth curve of a compound.
#' It identifies the row letter corresponding to the compound of interest, gathers water wells from row H and the end of the row containing the compound,
#' selects the DMSO well with the concentration used in compound dilution, and combines water and DMSO data into one dataframe.
#'
#' @param cname A character specifying the name of the compound for which baseline data is to be gathered.
#' @param conc_i The concentration of the compound for which baseline data is to be gathered.
#' @param growth_df A dataframe containing growth data for wells.
#' @param info_df A dataframe containing information about wells, such as compound and concentration.
#'
#' @return A dataframe representing the baseline data, including water and DMSO wells.
#'
#' @examples
#' gather_baseline("compound_name", 0.1, growth_data, well_info)
#'
# We gather the row letter that corresponds to our compound of interest.
compound_row <- growth_df %>%
mutate(row_letter = str_split(well, "", simplify = TRUE)[,1]) %>%
select(row_letter, compound) %>%
distinct() %>%
filter(compound == cname) %>%
select(row_letter) %>%
unlist()
# Now we can gather the Water wells corresponding to row H and the water well at the end of the row of our compound of interest
water_data <- growth_df %>%
mutate(row_letter = str_split(well, "", simplify = TRUE)[,1]) %>%
filter(compound == "Water", row_letter %in% c("H", compound_row)) %>%
select(-row_letter)
# We should select the DMSO well with the concentration of DMSO used in the compound dilution
dmso_perc <- well_metadata %>%
filter(compound == cname, concentration == conc_i) %>%
select(dmso_percentage) %>%
unlist()
# In a situation were no DMSO was used, we can just show the lowest concentration
if(dmso_perc == 0){
dmso_perc = 0.002
}
# Now we gather the corresponding well
dmso_well <- info_df %>%
filter(compound == "DMSO", concentration == dmso_perc) %>%
select(well) %>%
unlist()
dmso_data <- growth_df %>%
filter(well==dmso_well)
# Now that we have DMSO and Water, we combine them into one dataframe
baseline_data <- bind_rows(water_data, dmso_data)
return(baseline_data)
}
## Gather average OD measurements
gather_mean_data <- function(input_str, chem_data = complete_od, meta_df=well_metadata, deliver="mean_od"){
#' Gather mean data for a specific combination of organism, compound, and concentration.
#'
#' This function retrieves mean data for a given combination of organism, compound, and concentration from the complete optical density measurements dataframe.
#' Depending on the 'deliver' parameter, it can return either the complete optical density data for the specified combination, the mean growth curve, or both.
#'
#' @param input_str A character string representing the combination of organism, compound, and concentration (e.g., "organism-compound-concentration").
#' @param chem_data A dataframe containing complete optical density measurements. Defaults to complete_od.
#' @param meta_df A dataframe containing metadata information about wells. Defaults to well_metadata.
#' @param deliver A character specifying what to deliver: "mean_od" for mean growth curve, "allcurve_od" for complete optical density data,
#' and "both" for both mean growth curve and complete optical density data. Defaults to "mean_od".
#'
#' @return Depending on the 'deliver' parameter, it returns either the mean growth curve (dataframe), the all optical density data (dataframe), or both (list of dataframes).
#'
#' @examples
#' gather_mean_data("SA-Ebastine-128")
# Extract relevant combos
split_data <- str_split(input_str, "-", simplify = TRUE)
org_str <- split_data[,1]
compound_i <- split_data[,2]
concentration_i <- as.numeric(split_data[,3])
# First, we gather the data for our organism of interest
od_organism <- chem_data %>%
filter(organism == org_str)
# Now we gather the correct baseline data
baseline_od <- gather_baseline(compound_i, concentration_i, od_organism, meta_df)
# Now we gather the data for our compound - concentration of interest
chem_od <- od_organism %>%
filter(compound == compound_i, concentration == concentration_i) %>%
mutate(compound = factor(compound, levels = c("Water", "DMSO", compound_i)))
# Join everthing
allcurves_od <- bind_rows(baseline_od, chem_od)
if(deliver == "allcurve_od"){
return(allcurves_od)
}
# Now we can gather the mean growth curve of each compound
meancurves_od <- allcurves_od %>%
group_by(compound, timepoint) %>%
summarise(mean_od = mean(od), stdv = sd(od)) %>%
mutate(curve_id = compound) %>%
ungroup() %>%
mutate(organism = org_str,
compound = compound_i,
concentration = concentration_i)
if(deliver == "mean_od"){
return(meancurves_od)
}
if(deliver == "both"){
return(list("allcurve_od"=allcurves_od, "meancurve_od"=meancurves_od))
}
}
gather_panel <- function(compound_name, meta_df=well_metadata, chem_od = complete_od){
#' Gather data for a given compound from a panel of organisms with associated concentrations.
#'
#' This function takes a compound name and retrieves relevant data from the chemical optical density measurements dataframe.
#' Finally, it gathers mean data for all organism-concentration-compound combinations and returns the complete panel.
#'
#' @param compound_name A character specifying the name of the compound for which data is to be gathered.
#' @param meta_df A dataframe containing metadata information about wells. Defaults to well_metadata.
#' @param chem_od A dataframe containing chemical optical density measurements. Defaults to complete_od.
#'
#' @return A dataframe representing the complete panel of organism-concentration-compound combinations.
#'
#' @examples
#' gather_panel("Visomitin")
# Gather the relevant data for a given compound
all_chem_combinations <- chem_od %>%
filter(compound == compound_name) %>%
select(organism, compound, concentration) %>%
distinct() %>%
# Create a string with all info
unite(col = combo_str, organism, compound, concentration, sep="-") %>%
select(combo_str) %>%
unlist()
# Gather the data for all organism-concentration-compound combinations
complete_panel <- bind_rows(lapply(all_chem_combinations, gather_mean_data))
return(complete_panel)
}
# Gather the average amount of time spent for each timepoint
time_df <- complete_od %>%
select(timepoint, time_spent_h) %>%
distinct() %>%
group_by(timepoint) %>%
summarise(time_spent_avg = mean(time_spent_h))
Visomitin
viso.panel.data <- gather_panel("Visomitin") %>%
left_join(time_df, by="timepoint") %>%
mutate(organism = case_when(organism == "ECiAi" ~ "EC IAI1",
organism == "ECUTI" ~ "EC UTI",
TRUE ~ organism)) %>%
filter(curve_id != "Water")
viso.panel <- ggplot(viso.panel.data, aes(x=time_spent_avg, y=mean_od, ymin = mean_od - stdv, ymax = mean_od + stdv, color=curve_id)) +
geom_line() +
geom_pointrange(size=0.1, alpha=0.5) +
facet_grid(concentration~organism) +
scale_color_manual(values=c("#1f78b4", "#e7298a"),
breaks = c("DMSO", "Visomitin")) +
theme_light() +
theme(legend.position = "bottom",
axis.text = element_text(size = 5)) +
scale_x_continuous(breaks = 1:12) +
labs(x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound")
viso.panel
ggsave(plot=viso.panel, filename = file.path(OUTPUT_PANEL_PLOTS_DIR, "visomitin_panel.pdf"), width = 21, height = 25, units = "cm")
Elvitegravir.
elvi.panel.data <- gather_panel("Elvitegravir") %>%
left_join(time_df, by="timepoint") %>%
mutate(organism = case_when(organism == "ECiAi" ~ "EC IAI1",
organism == "ECUTI" ~ "EC UTI",
TRUE ~ organism)) %>%
filter(curve_id != "Water")
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
elvi.panel <- ggplot(elvi.panel.data, aes(x=time_spent_avg, y=mean_od, ymin = mean_od - stdv, ymax = mean_od + stdv, color=curve_id)) +
geom_line() +
geom_pointrange(size=0.1, alpha=0.5) +
facet_grid(concentration~organism) +
scale_color_manual(values=c("#1f78b4", "#e7298a"),
breaks = c("DMSO", "Elvitegravir")) +
theme_light() +
theme(legend.position = "bottom",
axis.text = element_text(size = 5)) +
scale_x_continuous(breaks = 1:12) +
labs(x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound")
elvi.panel
ggsave(plot=elvi.panel, filename = file.path(OUTPUT_PANEL_PLOTS_DIR, "elvitegravir_panel.pdf"), width = 21, height = 25, units = "cm")
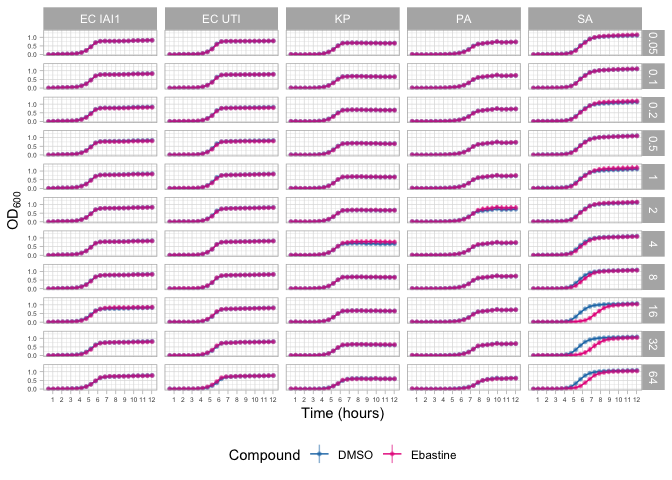
Ebastine.
ebas.panel.data <- gather_panel("Ebastine") %>%
left_join(time_df, by="timepoint") %>%
mutate(organism = case_when(organism == "ECiAi" ~ "EC IAI1",
organism == "ECUTI" ~ "EC UTI",
TRUE ~ organism)) %>%
filter(curve_id != "Water")
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
ebas.panel <- ggplot(ebas.panel.data, aes(x=time_spent_avg, y=mean_od, ymin = mean_od - stdv, ymax = mean_od + stdv, color=curve_id)) +
geom_line() +
geom_pointrange(size=0.1, alpha=0.5) +
facet_grid(concentration~organism) +
scale_color_manual(values=c("#1f78b4", "#e7298a"),
breaks = c("DMSO", "Ebastine")) +
theme_light() +
theme(legend.position = "bottom",
axis.text = element_text(size = 5)) +
scale_x_continuous(breaks = 1:12) +
labs(x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound")
ebas.panel
ggsave(plot=ebas.panel, filename = file.path(OUTPUT_PANEL_PLOTS_DIR, "ebastine_panel.pdf"), width = 21, height = 25, units = "cm")
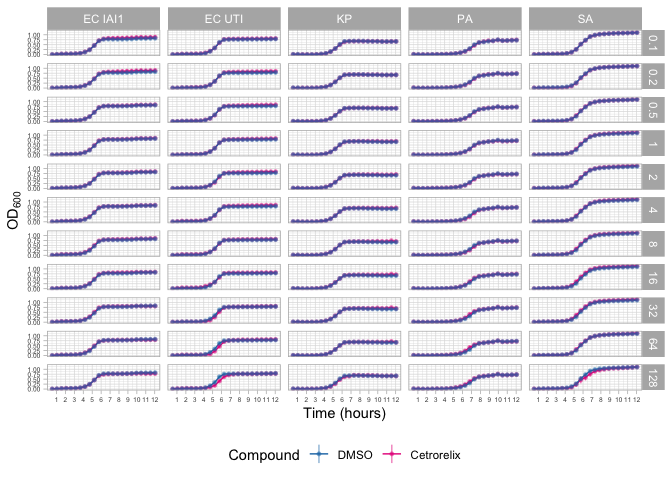
Cetrorelix
cetro.panel.data <- gather_panel("Cetrorelix") %>%
left_join(time_df, by="timepoint") %>%
mutate(organism = case_when(organism == "ECiAi" ~ "EC IAI1",
organism == "ECUTI" ~ "EC UTI",
TRUE ~ organism)) %>%
filter(curve_id != "Water")
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
cetro.panel <- ggplot(cetro.panel.data, aes(x=time_spent_avg, y=mean_od, ymin = mean_od - stdv, ymax = mean_od + stdv, color=curve_id)) +
geom_line() +
geom_pointrange(size=0.1, alpha=0.5) +
facet_grid(concentration~organism) +
scale_color_manual(values=c("#1f78b4", "#e7298a"),
breaks = c("DMSO", "Cetrorelix")) +
theme_light() +
theme(legend.position = "bottom",
axis.text = element_text(size = 5)) +
scale_x_continuous(breaks = 1:12) +
labs(x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound")
cetro.panel
ggsave(plot=cetro.panel, filename = file.path(OUTPUT_PANEL_PLOTS_DIR, "cetrorelix_panel.pdf"), width = 21, height = 25, units = "cm")
Opicapone
opica.panel.data <- gather_panel("Opicapone") %>%
left_join(time_df, by="timepoint") %>%
mutate(organism = case_when(organism == "ECiAi" ~ "EC IAI1",
organism == "ECUTI" ~ "EC UTI",
TRUE ~ organism)) %>%
filter(curve_id != "Water")
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
opica.panel <- ggplot(opica.panel.data, aes(x=time_spent_avg, y=mean_od, ymin = mean_od - stdv, ymax = mean_od + stdv, color=curve_id)) +
geom_line() +
geom_pointrange(size=0.1, alpha=0.5) +
facet_grid(concentration~organism) +
scale_color_manual(values=c("#1f78b4", "#e7298a"),
breaks = c("DMSO", "Opicapone")) +
theme_light() +
theme(legend.position = "bottom",
axis.text = element_text(size = 5)) +
scale_x_continuous(breaks = 1:12) +
labs(x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound")
opica.panel
ggsave(plot=opica.panel, filename = file.path(OUTPUT_PANEL_PLOTS_DIR, "opicapone_panel.pdf"), width = 21, height = 25, units = "cm")
Thymidine
thym.panel.data <- gather_panel("Thymidine") %>%
left_join(time_df, by="timepoint") %>%
mutate(organism = case_when(organism == "ECiAi" ~ "EC IAI1",
organism == "ECUTI" ~ "EC UTI",
TRUE ~ organism)) %>%
filter(curve_id != "Water")
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
## `summarise()` has grouped output by 'compound'. You can override using the
## `.groups` argument.
thym.panel <- ggplot(thym.panel.data, aes(x=time_spent_avg, y=mean_od, ymin = mean_od - stdv, ymax = mean_od + stdv, color=curve_id)) +
geom_line() +
geom_pointrange(size=0.1, alpha=0.5) +
facet_grid(concentration~organism) +
scale_color_manual(values=c("#1f78b4", "#e7298a"),
breaks = c("DMSO", "Thymidine")) +
theme_light() +
theme(legend.position = "bottom",
axis.text = element_text(size = 5)) +
scale_x_continuous(breaks = 1:12) +
labs(x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound")
thym.panel
ggsave(plot=thym.panel, filename = file.path(OUTPUT_PANEL_PLOTS_DIR, "thymidine_panel.pdf"), width = 21, height = 25, units = "cm")
MIC panel.
Here we look at the dose-response data for each compound.
gather_od <- function(df, t_i){
#' Gather optical density data at a specified timepoint.
#'
#' This function gathers optical density data at a specified timepoint for each strain replicate.
#' It determines the timepoint closest to the specified time of interest for each strain replicate,
#' then gathers the optical density data corresponding to that timepoint.
#'
#' @param df A dataframe containing optical density measurements.
#' @param t_i Numeric value representing the timepoint of interest.
#'
#' @return A dataframe containing optical density data at the specified timepoint.
#'
#' @examples
#' gather_od(df, 6)
#'
# Determine for each strain_rep the timepoint closest to our time of interest
tps_interest <- df %>%
group_by(strain_rep) %>%
# Timepoint - timespent relationship
select(strain_rep, timepoint, time_spent_h) %>%
distinct() %>%
# Gather the timepoint closest to our time of interest
mutate(diff_time = abs(t_i - time_spent_h)) %>%
slice_min(diff_time, n=1) %>%
select(strain_rep, timepoint) %>%
rename("tp_interest" = "timepoint")
# Gather relevant OD data
od_interest <- df %>%
left_join(tps_interest, by="strain_rep") %>%
filter(timepoint == tp_interest)
return(od_interest)
}
average_od <- function(df){
#' Calculate average optical density measurements per compound per concentration.
#'
#' This function calculates the average optical density measurements per compound per concentration.
#' It creates identifiers for each experiment, calculates the mean, median, and standard deviation of optical density measurements,
#' separates the experiment information into organism, compound, and concentration columns, and converts concentration to numeric and factor.
#'
#' @param df A dataframe containing optical density measurements.
#'
#' @return A dataframe containing average optical density measurements per compound per concentration.
#'
#' @details This function requires the tidyverse package to be installed and loaded in the R environment.
#'
#' @examples
#' average_od(df)
# Get the average OD per compound per concentration
od_avergaes_dose <- df %>%
# Create identifiers
mutate(experiment_info = paste(organism, compound, concentration, sep = "_")) %>%
# Gather mean per experiment
group_by(experiment_info) %>%
summarise(n_samples = n(), avg_od = mean(od), med_od = median(od), stdv = sd(od)) %>%
ungroup() %>%
# Un group
separate(experiment_info, into=c("organism", "compound", "concentration"), sep="_") %>%
mutate(concentration = as.numeric(concentration),
concentration = factor(concentration, levels=sort(unique(concentration))))
return(od_avergaes_dose)
}
dose_growth <- function(od_at_h, all_info=complete_od,
exclude_water = TRUE, return_all_od = FALSE){
#' Calculate dose-response growth curves based on optical density measurements.
#'
#' This function calculates dose-response growth curves based on optical density measurements at a specific timepoint.
#' It first creates a column containing the strain replicate information, gathers optical density at the specified timepoint,
#' calculates the average response for each compound at the specified timepoint, excludes water data if specified, and reorders compounds for plotting.
#'
#' @param od_at_h Numeric value representing optical density at the timepoint of interest.
#' @param all_info A dataframe containing all optical density measurements. Defaults to complete_od.
#' @param exclude_water Logical indicating whether to exclude water data from the analysis. Defaults to TRUE.
#'
#' @return A dataframe containing dose-response growth curves based on optical density measurements.
#'
#' @examples
#' dose_growth(0.5, complete_od)
# Create a column that contains the strain replicate
info_strainrep <- all_info %>%
mutate(strain_rep = paste(organism, replicate, sep="_"))
# Gather od at the time of interest
od_interest <- gather_od(info_strainrep, od_at_h)
if(return_all_od){
return(od_interest)
}
# Gather the average response at tp
od_averages <- average_od(od_interest)
# Exclude water
if(exclude_water){
od_averages <- od_averages %>%
filter(compound != "Water")
}
# Re-oder compounds
chemicals <- unique(od_averages$compound)
chemicals_wo_dmso <- sort(chemicals[chemicals!="DMSO"])
chemical_levels <- c("DMSO", chemicals_wo_dmso)
od_averages <- od_averages %>%
mutate(compound = factor(compound, levels=chemical_levels),
od_at_t = od_at_h)
return(od_averages)
}
OD after 10 hours of growth.
dose_response_10 <- dose_growth(10)
dose_reponse_10.p <- ggplot(dose_response_10, aes(x=concentration, y=avg_od,
ymin=avg_od - stdv, ymax=avg_od + stdv,
group=organism, color=organism)) +
geom_line() +
geom_pointrange(size=0.35, alpha=0.5) +
scale_color_manual(values=c("#1b9e77", "#d95f02", "#7570b3", "#e6ab02", "#e7298a"),
labels=c(expression(paste(italic("E. coli "), "IAI1")),
expression(paste(italic("E. coli "), "UTI")),
expression(italic("K. pneumoniae")),
expression(italic("P. aeruginosa")),
expression(italic("S. aureus")))) +
facet_wrap(~compound, ncol=4, scales="free_x") +
theme_bw() +
theme(legend.position = "right",
legend.text.align = 0,
text = element_text(size=10)) +
labs(x=latex2exp::TeX("Concentration ($\\mu$g/mL)"),
color="Organism",
y=latex2exp::TeX("$OD_{600}$ after 10 hours"))
dose_reponse_10.p
dose_response_10_complete <- dose_growth(10, return_all_od = TRUE)
dose_response_10_complete %>%
select(compound, concentration, organism, well, replicate, od, time_spent_h, curve_id) %>%
arrange(compound, concentration, organism) %>%
write_tsv(file.path(OUTPUT_DIR, "MIC_10h.tsv"))
Specific growth curves.
Here we look at specific growth curves.
Ebastine.
Ebastine effects on the growth of S. aureus.
# Gather the relevant data
ebas_od <- complete_od %>%
filter(organism == "SA",
compound %in% c("Ebastine", "DMSO", "Water"),
timepoint >= 2) %>%
mutate(chem_conc = paste(compound, concentration, sep="_")) %>%
filter(chem_conc %in% c("Ebastine_16", "Ebastine_32", "DMSO_0.64", "Water_0"),
str_detect(well, "^[HGD]"))
sa_ebastine_od <- ggplot() +
geom_line(data=subset(ebas_od, chem_conc == "Water_0"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=0.45) +
geom_line(data=subset(ebas_od, chem_conc == "DMSO_0.64"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=1) +
geom_line(data=subset(ebas_od, chem_conc == "Ebastine_16"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=1) +
#geom_line(data=subset(ebas_od, chem_conc == "Ebastine_32"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=1) +
scale_color_manual(values=c("grey", "#1f78b4", "#e7298a"),
labels=c("Water", latex2exp::TeX("DMSO (0.64 $\\mu$g/mL)"), latex2exp::TeX("Ebastine (16 $\\mu$g/mL)")), #, latex2exp::TeX("Ebastine (32 $\\mu$g/mL)")),
breaks = c("Water_0", "DMSO_0.64", "Ebastine_16")) + #, "Ebastine_32")) +
scale_x_continuous(breaks = 1:12) +
theme_bw() +
labs(title = expression(paste(italic("S. aureus "), "- Ebastine")),
x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound") +
theme(text = element_text(size=10))
sa_ebastine_od
ebas_od %>%
select(-timepoint) %>%
filter(chem_conc %in% c("Ebastine_16","DMSO_0.64", "Water_0"),
str_detect(well, "^[HGD]")) %>%
select(compound, concentration, organism, well, replicate, time_spent_h, od, dmso_percentage, curve_id) %>%
arrange(compound) %>%
write_tsv(file.path(OUTPUT_DIR, "ebastine_od.tsv"))
Opicapone.
Opicapone effects on the growth of S. aureus.
opi_od <- complete_od %>%
filter(organism == "SA",
compound %in% c("Opicapone", "DMSO", "Water"),
timepoint >= 2) %>%
mutate(chem_conc = paste(compound, concentration, sep="_")) %>%
filter(chem_conc %in% c("Opicapone_16", "Opicapone_128", "DMSO_2.56", "Water_0"),
str_detect(well, "^[HGB]"))
sa_opicapone_od <- ggplot() +
geom_line(data=subset(opi_od, chem_conc == "Water_0"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=0.45) +
geom_line(data=subset(opi_od, chem_conc == "DMSO_2.56"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=1) +
geom_line(data=subset(opi_od, chem_conc == "Opicapone_16"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=1) +
geom_line(data=subset(opi_od, chem_conc == "Opicapone_128"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=1) +
scale_color_manual(values=c("grey", "#1f78b4", "#fbb4b9", "#ae017e"),
labels=c("Water", latex2exp::TeX("DMSO (2.56 $\\mu$g/mL)"), latex2exp::TeX("Opicapone (16 $\\mu$g/mL)"), latex2exp::TeX("Opicapone (128 $\\mu$g/mL)")),
breaks = c("Water_0", "DMSO_2.56", "Opicapone_16", "Opicapone_128")) +
scale_x_continuous(breaks = 1:12) +
theme_bw() +
labs(title = expression(paste(italic("S. aureus "), "- Opicapone")),
x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound") +
theme(text = element_text(size=10),
legend.position = c(0.19, 0.77))
sa_opicapone_od
opi_od %>%
select(compound, concentration, organism, well, replicate, time_spent_h, od, dmso_percentage, curve_id) %>%
arrange(compound) %>%
write_tsv(file.path(OUTPUT_DIR, "opicapone_od.tsv"))
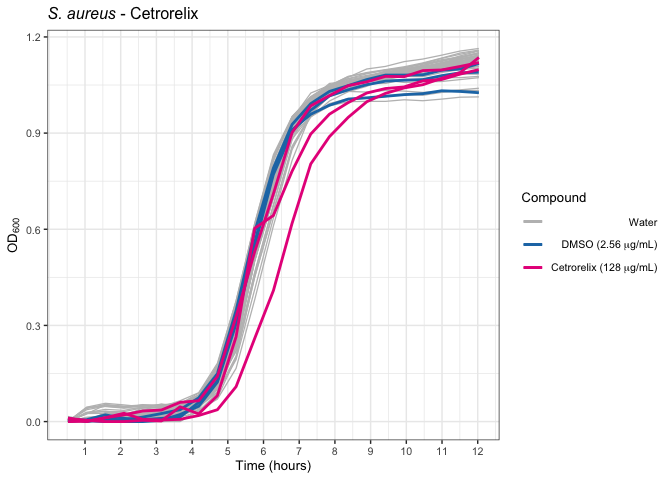
Cetrorelix
Ebastine effects on the growth of S. aureus.
# Gather the relevant data
cetro_od <- complete_od %>%
filter(organism == "SA",
compound %in% c("Cetrorelix", "DMSO", "Water"),
timepoint >= 2) %>%
mutate(chem_conc = paste(compound, concentration, sep="_")) %>%
filter(chem_conc %in% c("Cetrorelix_128", "DMSO_2.56", "Water_0"),
str_detect(well, "^[HGA]"))
sa_cetrorelix_od <- ggplot() +
geom_line(data=subset(cetro_od, chem_conc == "Water_0"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=0.45) +
geom_line(data=subset(cetro_od, chem_conc == "DMSO_2.56"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=1) +
geom_line(data=subset(cetro_od, chem_conc == "Cetrorelix_128"), aes(x=time_spent_h, y=od, group=curve_id, color=chem_conc), linewidth=1) +
scale_color_manual(values=c("grey", "#1f78b4", "#e7298a"),
labels=c("Water", latex2exp::TeX("DMSO (2.56 $\\mu$g/mL)"), latex2exp::TeX("Cetrorelix (128 $\\mu$g/mL)")),
breaks = c("Water_0", "DMSO_2.56", "Cetrorelix_128")) +
scale_x_continuous(breaks = 1:12) +
theme_bw() +
labs(title = expression(paste(italic("S. aureus "), "- Cetrorelix")),
x="Time (hours)",
y=latex2exp::TeX("$OD_{600}$"),
color="Compound") +
theme(text = element_text(size=10))
sa_cetrorelix_od
Sigmoid Modelling.
library(sicegar)
# Function to check growth
checkGrowth <- function(growth_df, threshold=0.2, n_above_threshold=2){
# Check if at least n_above_threshold timepoints pass the OD threshold
maxOD_check <- sum(growth_df$intensity > threshold) >= n_above_threshold
return(maxOD_check)
}
# Function to calculate the fitted curve
logistic_parametrized <- function(x, Imax, xmid, alpha){
y <- Imax / (1 + exp(-alpha*(x-xmid)))
return(y)
}
# Global sigmoid fitting function
fit_sigmoids <- function(cid, data_df, base_output=NULL, minimum_od = 0.2, n_above_minOD=2){
#' This function performs growth check, normalisation, sigmoid fitting and parameter estimation for a given growth curve.
#' @param curve_id some identifier that allows for the selection of an individual curve form data_df
#' @param data_df Data.Frame that contains all of the growth information
#' @param base_output data.frame that acts as placeholder for unsuccessful growth curve fits
#' @param minimum_od value that indicates the OD threshold that must be surpassed by a growth in order to be considered "alive"
#' @param n_above_minOD value that indicates the number of timepoints where the OD value is greater than minimum_od
#' @return data.frame contains information on whether the curve passed the growth pre-check and, if so, the parameters of the estimated logistic curve.
# Get the relevant data
curve_od <- data_df %>%
filter(curve_id == cid) %>%
select(time_spent_h, od) %>%
rename("time" = "time_spent_h", "intensity" = "od")
# Check if the curve is growing
precheck <- checkGrowth(growth_df = curve_od, threshold = minimum_od, n_above_threshold = n_above_minOD)
# Output if the check fails
if (precheck == FALSE){
out_df <- base_output
out_df$dataInputName <- cid
out_df$isThisaFit <- FALSE
out_df$model <- "no_model"
out_df$additionalParameters <- FALSE
out_df$maximum_x <- NA
return(out_df)
}
# If there is a signal, fit the data
# Normalise the data
curve_od.norm <- sicegar::normalizeData(dataInput = curve_od, dataInputName = cid)
# Perform modelling
set.seed(42)
sigmoidalModel <- multipleFitFunction(dataInput=curve_od.norm,
model="sigmoidal")
# Additional Parameters
fit_vector <- parameterCalculation(sigmoidalModel)
# Format as datafrane
out_df <- data.frame(fit_vector)
return(out_df)
}
example_out <- fit_sigmoids(cid = "ECiAi_F1_1", data_df = complete_od)
for(c in colnames(example_out)){
example_out[, c] <- 0
}
Apply sigmoid function to all curves.
library(foreach) # Parallelization
##
## Attaching package: 'foreach'
## The following objects are masked from 'package:purrr':
##
## accumulate, when
doParallel::registerDoParallel(cores=10)
fit.curves <- foreach(cid = unique(complete_od$curve_id), .combine = "rbind") %dopar% fit_sigmoids(cid, complete_od, example_out)
Add generation time.
growth.params <- fit.curves %>%
# Relevant growth parameters
select(dataInputName, startPoint_x, reachMaximum_x, maximum_y, midPoint_x, slopeParam_Estimate) %>%
separate(dataInputName, into = c("organism", "well", "replicate"), sep="_", remove = FALSE) %>%
# Add generation time
mutate(generation_time = log(2) / slopeParam_Estimate,
generation_time = if_else(generation_time == Inf, 0, generation_time)) %>%
# Add metadata
left_join(well_metadata, by="well")
Compare growth parameters
GPARAM_OUTPUT_DIR <- file.path(OUTPUT_DIR, "growth_parameter_plots")
if(!dir.exists(GPARAM_OUTPUT_DIR)){
dir.create(GPARAM_OUTPUT_DIR)
}
# Gather the growth parameters
gather_gparam <- function(compound_name, gparam_df = growth.params, meta_df = well_metadata){
#' Gather growth parameters for a specific compound.
#'
#' This function gathers growth parameters for a specific compound from a growth parameter dataframe.
#' The resulting dataframe includes parameters for the specified compound, DMSO, and water.
#'
#' @param compound_name A character specifying the name of the compound for which parameters are to be gathered.
#' @param gparam_df A dataframe containing growth parameters. Defaults to growth.params.
#' @param meta_df A dataframe containing metadata information about wells. Defaults to well_metadata.
#'
#' @return A dataframe containing growth parameters for the specified compound, DMSO, and water.
#'
#' @examples
#' gather_gparam("compound_name", growth.params, well_metadata)
# Gather compound parameters
compound.parameters <- gparam_df %>%
filter(compound == compound_name) %>%
# Add column for chemical concentration
mutate(chem_conc = concentration)
# Gather the corresponding DMSO percentage
conc.map <- compound.parameters %>%
select(concentration, dmso_percentage) %>%
distinct() %>%
rename("chem_conc" = "concentration")
# If DMSO == 0 add the common concentration range
if(all(conc.map$dmso_percentage == 0)){
conc.map <- gparam_df %>%
filter(compound == "Opicapone") %>%
select(concentration, dmso_percentage) %>%
distinct() %>%
rename("chem_conc" = "concentration")
}
# Gather the DMSO well
dmso.params <- gparam_df %>%
filter(compound %in% c("DMSO")) %>%
left_join(conc.map, by="dmso_percentage") %>%
filter(!is.na(chem_conc))
# Add water parameters
water_params <- gparam_df %>%
filter(compound %in% c("Water"), str_detect(well, "^H"))
complete_water <- bind_rows(lapply(unique(compound.parameters$chem_conc), function(x){water_params$chem_conc = x; water_params}))
# Combine all data
complete_params <- bind_rows(complete_water, dmso.params, compound.parameters) %>%
mutate(compound = factor(compound, levels = c("Water", "DMSO", compound_name)))
return(complete_params)
}
Visomitin.
# Specify compound
compound_n <- "Visomitin"
# Gather the parameters
compound.params <- gather_gparam(compound_n)
# Maximum OD plot
maxod.bp <- ggplot(compound.params, aes(x=compound, y=maximum_y, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Maximum OD"))
# End of Lag Phase
lagphase.bp <- ggplot(compound.params, aes(x=compound, y=startPoint_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- End of Lag Phase"))
# Start Stationary
statphase.bp <- ggplot(compound.params, aes(x=compound, y=reachMaximum_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Start Stationary Phase"))
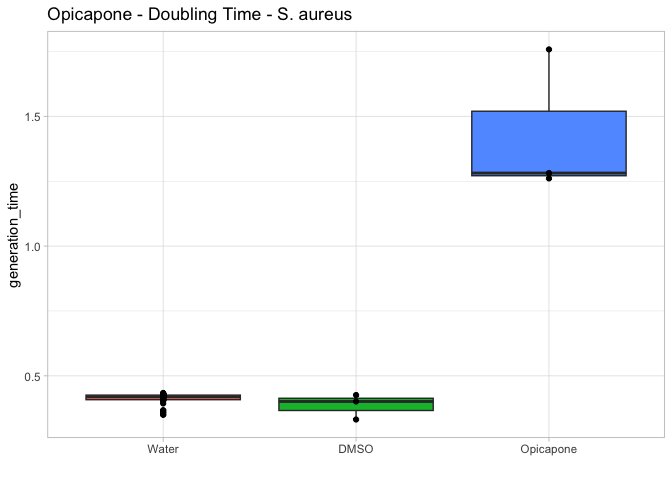
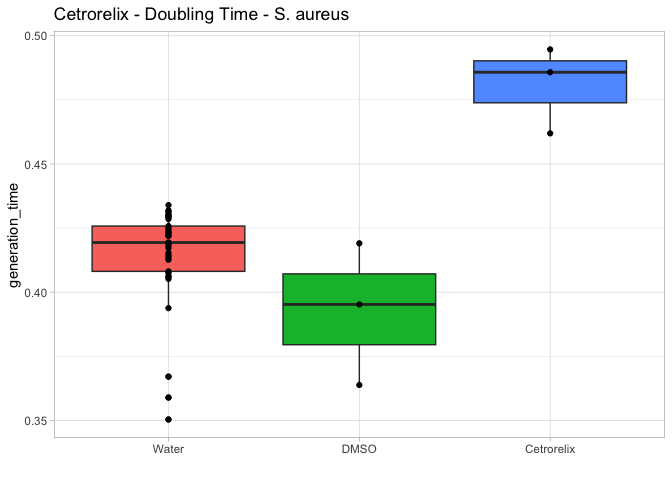
# Generation Time
gentime.bp <- ggplot(compound.params, aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Generation Time"))
ggsave(plot=maxod.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_maxOD.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=lagphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_lagPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=statphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_stationaryPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=gentime.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_genTime.pdf")), width = 21, height = 25, units = "cm")
Elvitegravir
# Specify compound
compound_n <- "Elvitegravir"
# Gather the parameters
compound.params <- gather_gparam(compound_n)
# Maximum OD plot
maxod.bp <- ggplot(compound.params, aes(x=compound, y=maximum_y, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Maximum OD"))
# End of Lag Phase
lagphase.bp <- ggplot(compound.params, aes(x=compound, y=startPoint_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- End of Lag Phase"))
# Start Stationary
statphase.bp <- ggplot(compound.params, aes(x=compound, y=reachMaximum_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Start Stationary Phase"))
# Generation Time
gentime.bp <- ggplot(compound.params, aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Generation Time"))
ggsave(plot=maxod.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_maxOD.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=lagphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_lagPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=statphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_stationaryPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=gentime.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_genTime.pdf")), width = 21, height = 25, units = "cm")
Ebastine
# Specify compound
compound_n <- "Ebastine"
# Gather the parameters
compound.params <- gather_gparam(compound_n)
# Maximum OD plot
maxod.bp <- ggplot(compound.params, aes(x=compound, y=maximum_y, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Maximum OD"))
# End of Lag Phase
lagphase.bp <- ggplot(compound.params, aes(x=compound, y=startPoint_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- End of Lag Phase"))
# Start Stationary
statphase.bp <- ggplot(compound.params, aes(x=compound, y=reachMaximum_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Start Stationary Phase"))
# Generation Time
gentime.bp <- ggplot(compound.params, aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Generation Time"))
ggsave(plot=maxod.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_maxOD.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=lagphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_lagPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=statphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_stationaryPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=gentime.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_genTime.pdf")), width = 21, height = 25, units = "cm")
library(ggpubr)
##
## Attaching package: 'ggpubr'
## The following object is masked from 'package:ggtree':
##
## rotate
sa.ebas.lg <- compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=startPoint_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
scale_fill_manual(values=c("grey", "#1f78b4", "#DE1F84")) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10)) +
labs(x="",
y="Time of end of lag-phase (h)",
title = "Time of end of lag-phase")
sa.ebas.sp <- compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=reachMaximum_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
scale_fill_manual(values=c("grey", "#1f78b4", "#DE1F84")) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10)) +
labs(x="",
y="Time of start of stationary-phase (h)",
title = "Time of start of stationary-phase")
sa.ebas.maxOD <- compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=maximum_y, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
scale_fill_manual(values=c("grey", "#1f78b4", "#DE1F84")) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10)) +
labs(x="",
y="Maximum OD",
title = "Maximum amount of growth")
sa.ebas.gent <- compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
scale_fill_manual(values=c("grey", "#1f78b4", "#DE1F84")) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10)) +
labs(x="",
y="Doubling time (h)",
title = "Greatest Doubling Time")
fig_growth = ggarrange(sa.ebas.maxOD, sa.ebas.gent,
sa.ebas.lg, sa.ebas.sp,
nrow = 2,
ncol = 2,
labels = c("a", "b", "c", "d"),
font.label = list(size = 12, color = "black", face = "plain"))
fig_growth
ggsave(plot=fig_growth, filename = file.path(OUTPUT_DIR, "sa_ebastine_gparam_boxplots.pdf"), dpi=300, height = 15, width = 21, units = "cm")
We can look at the average increase in the start of the growth
compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
group_by(compound) %>%
summarise(average_lp = mean(startPoint_x), stdv_lp = sd(startPoint_x)) %>%
knitr::kable()
| compound | average_lp | stdv_lp |
|---|---|---|
| Water | 4.622365 | 0.1355105 |
| DMSO | 4.613593 | 0.0609431 |
| Ebastine | 6.337734 | 0.2433150 |
paste("Delay of onset of growth by", 6.337735 - 4.613593, "hours")
## [1] "Delay of onset of growth by 1.724142 hours"
Opicapone
# Specify compound
compound_n <- "Opicapone"
# Gather the parameters
compound.params <- gather_gparam(compound_n)
# Maximum OD plot
maxod.bp <- ggplot(compound.params, aes(x=compound, y=maximum_y, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Maximum OD"))
# End of Lag Phase
lagphase.bp <- ggplot(compound.params, aes(x=compound, y=startPoint_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- End of Lag Phase"))
# Start Stationary
statphase.bp <- ggplot(compound.params, aes(x=compound, y=reachMaximum_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Start Stationary Phase"))
# Generation Time
gentime.bp <- ggplot(compound.params, aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Generation Time"))
ggsave(plot=maxod.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_maxOD.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=lagphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_lagPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=statphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_stationaryPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=gentime.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_genTime.pdf")), width = 21, height = 25, units = "cm")
sa.opi.lg <- compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=startPoint_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
scale_fill_manual(values=c("grey", "#1f78b4", "#DE1F84")) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10)) +
labs(x="",
y="Time of end of lag-phase (h)",
title = "Time of end of lag-phase")
sa.opi.sp <- compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=reachMaximum_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
scale_fill_manual(values=c("grey", "#1f78b4", "#DE1F84")) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10)) +
labs(x="",
y="Time of start of stationary-phase (h)",
title = "Time of start of stationary-phase")
sa.opi.maxOD <- compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=maximum_y, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
scale_fill_manual(values=c("grey", "#1f78b4", "#DE1F84")) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10)) +
labs(x="",
y="Maximum OD",
title = "Maximum amount of growth")
sa.opi.gent <- compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
scale_fill_manual(values=c("grey", "#1f78b4", "#DE1F84")) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10)) +
labs(x="",
y="Doubling time (h)",
title = "Greatest Doubling Time")
fig_growth = ggarrange(sa.opi.maxOD, sa.opi.gent,
sa.opi.lg, sa.opi.sp,
nrow = 2,
ncol = 2,
labels = c("a", "b", "c", "d"),
font.label = list(size = 12, color = "black", face = "plain"))
fig_growth
ggsave(plot=fig_growth, filename = file.path(OUTPUT_DIR, "sa_opicapone_gparam_boxplots.pdf"), dpi=300, height = 15, width = 21, units = "cm")
compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
group_by(compound) %>%
summarise(average_lp = mean(startPoint_x), stdv_lp = sd(startPoint_x)) %>%
knitr::kable()
| compound | average_lp | stdv_lp |
|---|---|---|
| Water | 4.622365 | 0.1355105 |
| DMSO | 4.588387 | 0.0641173 |
| Opicapone | 5.659881 | 0.2791081 |
paste("Delay of onset of growth by", 5.659881 - 4.588387, "hours")
## [1] "Delay of onset of growth by 1.071494 hours"
compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
ggplot(aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
theme_light() +
theme(legend.position = "none") +
labs(x="",
title = paste(compound_n, "- Doubling Time - S. aureus"))
compound.params %>%
filter(organism == "SA",
chem_conc == "16") %>%
group_by(compound) %>%
summarise(average_gt = mean(generation_time), stdv_gt = sd(generation_time)) %>%
knitr::kable()
| compound | average_gt | stdv_gt |
|---|---|---|
| Water | 0.4141490 | 0.0202054 |
| DMSO | 0.3863526 | 0.0488593 |
| Opicapone | 1.4331948 | 0.2811447 |
Cetrorelix
# Specify compound
compound_n <- "Cetrorelix"
# Gather the parameters
compound.params <- gather_gparam(compound_n)
# Maximum OD plot
maxod.bp <- ggplot(compound.params, aes(x=compound, y=maximum_y, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Maximum OD"))
# End of Lag Phase
lagphase.bp <- ggplot(compound.params, aes(x=compound, y=startPoint_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- End of Lag Phase"))
# Start Stationary
statphase.bp <- ggplot(compound.params, aes(x=compound, y=reachMaximum_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Start Stationary Phase"))
# Generation Time
gentime.bp <- ggplot(compound.params, aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Generation Time"))
ggsave(plot=maxod.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_maxOD.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=lagphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_lagPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=statphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_stationaryPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=gentime.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_genTime.pdf")), width = 21, height = 25, units = "cm")
compound.params %>%
filter(organism == "SA",
chem_conc == "128") %>%
ggplot(aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
theme_light() +
theme(legend.position = "none") +
labs(x="",
title = paste(compound_n, "- Doubling Time - S. aureus"))
compound.params %>%
filter(organism == "SA",
chem_conc == "128") %>%
group_by(compound) %>%
summarise(average_gt = mean(generation_time), stdv_gt = sd(generation_time)) %>%
knitr::kable()
| compound | average_gt | stdv_gt |
|---|---|---|
| Water | 0.4141490 | 0.0202054 |
| DMSO | 0.3926798 | 0.0277075 |
| Cetrorelix | 0.4807707 | 0.0169240 |
Thymidine
# Specify compound
compound_n <- "Thymidine"
# Gather the parameters
compound.params <- gather_gparam(compound_n)
# Maximum OD plot
maxod.bp <- ggplot(compound.params, aes(x=compound, y=maximum_y, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Maximum OD"))
# End of Lag Phase
lagphase.bp <- ggplot(compound.params, aes(x=compound, y=startPoint_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- End of Lag Phase"))
# Start Stationary
statphase.bp <- ggplot(compound.params, aes(x=compound, y=reachMaximum_x, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Start Stationary Phase"))
# Generation Time
gentime.bp <- ggplot(compound.params, aes(x=compound, y=generation_time, fill=compound)) +
geom_boxplot(width=0.8) +
geom_point(fill="black") +
facet_grid(chem_conc~organism) +
theme_light() +
theme(legend.position = "none",
text = element_text(size=10),
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1.25)) +
labs(x="",
title = paste(compound_n, "- Generation Time"))
ggsave(plot=maxod.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_maxOD.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=lagphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_lagPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=statphase.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_stationaryPhase.pdf")), width = 21, height = 25, units = "cm")
ggsave(plot=gentime.bp, filename = file.path(GPARAM_OUTPUT_DIR, paste0(compound_n, "_genTime.pdf")), width = 21, height = 25, units = "cm")
Write parameters.
write_tsv(growth.params, file.path(OUTPUT_DIR, "growth_parameters.tsv.gz"))
Session Information
sessionInfo()
## R version 4.3.1 (2023-06-16)
## Platform: x86_64-apple-darwin20 (64-bit)
## Running under: macOS Ventura 13.4.1
##
## Matrix products: default
## BLAS: /Library/Frameworks/R.framework/Versions/4.3-x86_64/Resources/lib/libRblas.0.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/4.3-x86_64/Resources/lib/libRlapack.dylib; LAPACK version 3.11.0
##
## locale:
## [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
##
## time zone: Europe/Berlin
## tzcode source: internal
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] ggpubr_0.6.0 foreach_1.5.2 sicegar_0.2.4 ggtree_3.8.2
## [5] readxl_1.4.2 lubridate_1.9.2 forcats_1.0.0 stringr_1.5.0
## [9] dplyr_1.1.2 purrr_1.0.2 readr_2.1.4 tidyr_1.3.0
## [13] tibble_3.2.1 ggplot2_3.4.4 tidyverse_2.0.0
##
## loaded via a namespace (and not attached):
## [1] tidyselect_1.2.0 farver_2.1.1 fastmap_1.1.1 latex2exp_0.9.6
## [5] lazyeval_0.2.2 digest_0.6.33 timechange_0.2.0 lifecycle_1.0.4
## [9] tidytree_0.4.6 magrittr_2.0.3 compiler_4.3.1 rlang_1.1.4
## [13] tools_4.3.1 utf8_1.2.4 yaml_2.3.7 knitr_1.43
## [17] ggsignif_0.6.4 labeling_0.4.3 bit_4.0.5 RColorBrewer_1.1-3
## [21] aplot_0.2.3 abind_1.4-5 withr_2.5.2 grid_4.3.1
## [25] fansi_1.0.6 colorspace_2.1-0 scales_1.3.0 iterators_1.0.14
## [29] cli_3.6.3 rmarkdown_2.24 crayon_1.5.2 ragg_1.2.5
## [33] treeio_1.24.3 generics_0.1.3 rstudioapi_0.15.0 tzdb_0.4.0
## [37] ape_5.8 parallel_4.3.1 ggplotify_0.1.2 cellranger_1.1.0
## [41] yulab.utils_0.1.7 vctrs_0.6.5 jsonlite_1.8.7 carData_3.0-5
## [45] minpack.lm_1.2-3 car_3.1-2 gridGraphics_0.5-1 hms_1.1.3
## [49] patchwork_1.1.2 bit64_4.0.5 rstatix_0.7.2 systemfonts_1.0.4
## [53] glue_1.7.0 codetools_0.2-19 cowplot_1.1.1 stringi_1.7.12
## [57] gtable_0.3.4 munsell_0.5.0 pillar_1.9.0 htmltools_0.5.6
## [61] R6_2.5.1 textshaping_0.3.6 doParallel_1.0.17 vroom_1.6.3
## [65] evaluate_0.21 lattice_0.21-8 highr_0.10 backports_1.4.1
## [69] broom_1.0.5 ggfun_0.1.6 Rcpp_1.0.13 nlme_3.1-162
## [73] xfun_0.40 fs_1.6.3 pkgconfig_2.0.3