45 KiB
Analysing MCE predictions
Roberto Olayo Alarcon 27/03/2024
In this file we analyse the predictions made in the previous step. At this point a literature search has been performed in order to validate predictions
library(tidyverse)
library(ggrepel)
library(readxl)
library(uwot)
Prepare directories.
PREDICTIONS_DIR <- "../data/04.new_predictions"
OUTPUT_DIR <- "../data/05.analyze_mce_predictions"
# Create output dir
if(!dir.exists(OUTPUT_DIR)){
dir.create(OUTPUT_DIR)
}
Library Representation.
In this section, we perform dimensionality reduction on the MolE representation of the MedChemExpress library for which we make predictions.
# Read Prediction Output
mole_predictions <- read_excel(file.path(PREDICTIONS_DIR, "mole_mce_predictions_litsearch.xlsx"), sheet = "mole_prediction_overview")
mole_predictions.over10 <- read_excel(file.path(PREDICTIONS_DIR, "mole_mce_predictions_litsearch.xlsx"), sheet = "mole_over10") %>%
mutate(`Reported Activity` = if_else(is.na(`Reported Activity`), "None", `Reported Activity`))
# Read the representation
mole_representation <- vroom::vroom(file.path(PREDICTIONS_DIR, "MolE_representation_medchemexpress.tsv.gz"),
show_col_types = FALSE) %>%
# Rename id column
rename("Catalog Number" = "...1") %>%
# Filter for the molecules for which we make predictions
filter(`Catalog Number` %in% mole_predictions$`Catalog Number`) %>%
# Set column to row.names
column_to_rownames("Catalog Number")
## New names:
## • `` -> `...1`
# Read Prediction Output
ecfp4_predictions <- read_excel(file.path(PREDICTIONS_DIR, "ecfp4_mce_predictions_litsearch.xlsx"), sheet = "ecfp4_prediction_overview")
ecfp4_predictions.over10 <- read_excel(file.path(PREDICTIONS_DIR, "ecfp4_mce_predictions_litsearch.xlsx"), sheet = "ecfp4_over10") %>%
mutate(`Reported Activity` = if_else(is.na(`Reported Activity`), "None", `Reported Activity`))
# Read the representation
ecfp4_representation <- vroom::vroom(file.path(PREDICTIONS_DIR, "ecfp4_representation_medchemexpress.tsv.gz"),
show_col_types = FALSE) %>%
# Rename id column
rename("Catalog Number" = "...1") %>%
# Filter for the molecules for which we make predictions
filter(`Catalog Number` %in% mole_predictions$`Catalog Number`) %>%
# Set column to row.names
column_to_rownames("Catalog Number")
## New names:
## • `` -> `...1`
Preprocess representation.
Remove constant and correlated features
remove_constant <- function(m.df, var.cutoff){
#' Removes constant columns from a data frame based on a variance threshold.
#'
#' @param m.df A data frame. Input data frame.
#' @param var.cutoff A numeric. Variance threshold. Columns with variance below this threshold will be removed.
#'
#' @return A data frame. Data frame with constant columns removed.
#'
#' @examples
#' remove_constant(m.df = my_data_frame, var.cutoff = 0.01)
#'
#' The function calculates the variance of each dimension in the input data frame.
#' It then identifies columns with variance above the specified threshold and removes constant columns.
#' The resulting data frame without constant columns is returned.
#'
# Variance of each dimension
var.feats <- apply(m.df, 2, var)
# Keep columns above variance thresh
keep_cols <- names(var.feats[var.feats > var.cutoff])
# Filter df
m.var.df <- m.df %>%
select(all_of(as.character(keep_cols)))
return(m.var.df)
}
remove_corfeats <- function(m.df, cor.thresh){
#' Removes correlated features from a data frame based on a correlation threshold.
#'
#' @param m.df A data frame. Input data frame.
#' @param cor.thresh A numeric. Correlation threshold.
#'
#' @return A data frame. Data frame with correlated features removed.
#'
#' @examples
#' remove_corfeats(m.df = my_data_frame, cor.thresh = 0.7)
#'
#' The function calculates the correlation matrix of the input data frame.
#' It then identifies correlated columns based on the correlation threshold.
#' Columns with correlations above the threshold are removed.
#' The resulting data frame without correlated features is returned.
#'
# Correlation Matrix
cor.mat <- cor(m.df)
# Format triangle
cor.mat[lower.tri(cor.mat, diag = TRUE)] <- 0
# Find correlated columns
corr.descision <- apply(cor.mat, 2, function(x){ifelse(any(x >= cor.thresh), "remove", "remain")})
# Keep columns
keep_cols <- names(corr.descision[corr.descision == "remain"])
# Filter df
m.uncor <- m.df %>%
select(all_of(keep_cols))
return(m.uncor)
}
preprocess_mole <- function(mole_df, min.var, corr.thresh){
#' Preprocesses molecular data by filtering out constant columns and correlated features.
#'
#' @param mole_df A data frame. Input molecular data frame.
#' @param min.var A numeric. Variance threshold for removing constant columns.
#' @param corr.thresh A numeric. Correlation threshold for removing correlated features.
#'
#' @return A data frame. Preprocessed molecular data frame.
#'
#' @examples
#' preprocess_mole(mole_df = my_molecular_data_frame, min.var = 0.01, corr.thresh = 0.7)
#'
#' The function preprocesses molecular data by first removing constant columns using the specified variance threshold.
#' Then, it removes correlated features based on the correlation threshold.
#' The resulting data frame is the preprocessed molecular data with constant columns and correlated features removed.
#'
# Filter variance
mole.variable <- remove_constant(mole_df, min.var)
# Filter correlated
mole.decorrelated <- remove_corfeats(mole.variable, corr.thresh)
return(mole.decorrelated)
}
# Prepare data
mole_prepared <- preprocess_mole(mole_representation, min.var=0.01, corr.thresh=0.90)
mole_prepared %>% dim()
## [1] 2319 445
Similar procedure for ECFP4 representation
hamming <- function(X) {
D <- (1 - X) %*% t(X)
HamDist <- D + t(D)
return(HamDist / ncol(X))
}
remove_rare <- function(f.df, min_chems){
# Removes columns where a feature is not present in the majority of compounds
#
# Parameters
# ----------
# - fdf: pandas dataframe. Columns are bits of the fingerprint. Rows are compounds
# - mc: int. The minimum number of molecules a feature should be present in order for it to be preserved
# Number of chemicals a feature is present in
n.freq <- colSums(f.df)
names(n.freq) <- colnames(f.df)
# Filter popular features
common.features <- n.freq[n.freq >= min_chems]
# Alter df
f.common <- f.df %>%
select(all_of(names(common.features)))
return(f.common)
}
remove_correlated <- function(f.df, min_dist){
# Removes highly similar features
#
# Parameters
# ----------
# - fdf: pandas dataframe. Dataframe where columns are bits in the fingerprint
# - mdist: float. Is minimal distance between features in order to be different enough
#
# Returns
# -------
# pandas dataframe
# Highly correlated features are removed (one of them is preserved)
# Hamming distance
dist.matrix <- hamming(t(f.df))
# Format
dist.matrix[lower.tri(dist.matrix, diag = TRUE)] <- 1
# Find correlated columns
corr.descision <- apply(dist.matrix, 2, function(x){ifelse(any(x <= min_dist), "remove", "remain")})
# Only keep un-correlated feats
keep_cols <- names(corr.descision[corr.descision == "remain"])
# Filter df
f.uncorr <- f.df %>%
select(all_of(as.character(keep_cols)))
return(f.uncorr)}
preprocess_fps <- function(fps_df, min_compounds, sim_threshold){
# Processes a fingerprint dataframe. Removes rare and highly correlated features
#
# Parameters
# ----------
# - fps_df: pandas dataframe. Dataframe where columns are bits in the fingerprint
# - min_compounds: int. The minimum number of molecules a feature should be present in order for it to be preserved
# - sim_threshold: float. Is minimal distance between features in order to be different enough
fps_common <- remove_rare(fps_df, min_compounds)
fps.decorr <- remove_correlated(fps_common, sim_threshold)
return(fps.decorr)
}
ecfp4_prepared <- preprocess_fps(ecfp4_representation, 10, 0.01)
ecfp4_prepared %>% dim()
## [1] 2319 992
Peform UMAP reduction.
set.seed(1234)
mole.umap <- umap(X=mole_prepared,
n_neighbors = 25,
n_components = 2,
min_dist = 0.35,
n_threads = 20,
metric="cosine")
mole.umap <- as.data.frame(mole.umap) %>%
rownames_to_column("catalog_number")
mole.umap %>%
ggplot(aes(x=V1, y=V2)) +
geom_point(alpha=0.9, color="white", fill="#C5C5C5",
size=2, shape=21, stroke=0.2) +
theme_classic() +
labs(x="UMAP 1",
y = "UMAP 2",
title = "MolE representation of MCE")
# Jaccard distance
jac.dist <- dist(ecfp4_prepared, method = "binary")
set.seed(1234)
ecfp4.umap <- umap(X=jac.dist,
n_neighbors = 20,
n_components = 2,
min_dist = 0.3,
n_threads = 20)
ecfp4.umap <- as.data.frame(ecfp4.umap) %>%
rownames_to_column("catalog_number")
ecfp4.umap %>%
ggplot(aes(x=V1, y=V2)) +
geom_point(alpha=0.9, color="white", fill="#C5C5C5",
size=2, shape=21, stroke=0.2) +
theme_classic() +
labs(x="UMAP 1",
y = "UMAP 2",
title = "ECFP4 representation of MCE")
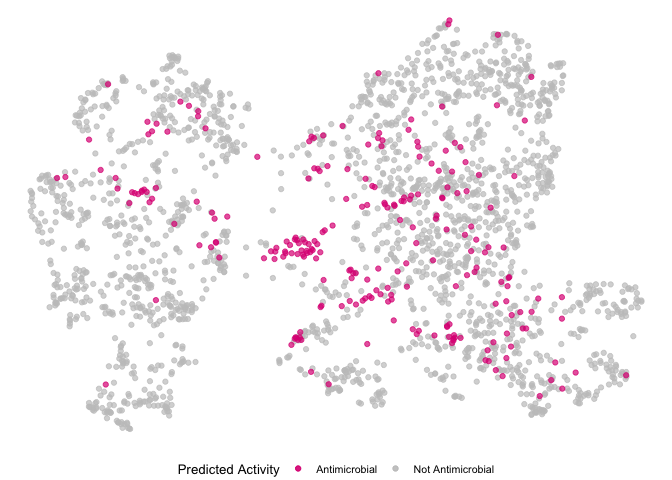
Plotting predicted broad-spectrum.
# Add information of broad-spectrum activity
mole.umap <- mole.umap %>%
mutate(pred_activity = if_else(catalog_number %in% mole_predictions.over10$`Catalog Number`, "Antimicrobial", "Not Antimicrobial"))
# Plot
m.umap.over10 <- ggplot(mole.umap, aes(x=V1, y=V2)) +
geom_point(data=subset(mole.umap, pred_activity == "Not Antimicrobial"), aes(color="Not Antimicrobial"), size=1.5) +
geom_point(data=subset(mole.umap, pred_activity == "Antimicrobial"), aes(color="Antimicrobial"), size=1.5) +
scale_color_manual(values=c("Antimicrobial" = alpha("#DE1F84", 0.7), "Not Antimicrobial" = alpha("#C5C5C5", 0.7))) +
theme_void() +
#coord_fixed(ratio = 0.4) +
theme(legend.position = "bottom",
axis.ticks.x = element_blank(),
axis.text.x = element_blank(),
axis.ticks.y = element_blank(),
axis.text.y = element_blank(),
text=element_text(size=10),
panel.background = element_rect(fill = "transparent", color=NA),
plot.background = element_rect(fill = "transparent", colour = NA)) +
labs(x="UMAP 1",
y="UMAP 2",
color="Predicted Activity")
m.umap.over10
chems_interst <- mole_predictions.over10 %>%
filter(ProductName %in% c("Ospemifene", "Shionone", "Bekanamycin", "Doxifluridine", "Ellagic acid")) %>%
select(`Catalog Number`, ProductName) %>%
rename("catalog_number" = "Catalog Number")
chems_interest_umap <- chems_interst %>%
left_join(mole.umap, by="catalog_number")
# Plot
m.umap.over10.annot <- ggplot(mole.umap, aes(x=V1, y=V2)) +
geom_point(data=subset(mole.umap, pred_activity == "Not Antimicrobial"), aes(color="Not Antimicrobial"), size=1.5) +
geom_point(data=subset(mole.umap, pred_activity == "Antimicrobial"), aes(color="Antimicrobial"), size=1.5) +
geom_text_repel(data = chems_interest_umap, aes(label=ProductName),
max.overlaps = Inf, size=2, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2, fontface="bold") +
scale_color_manual(values=c("Antimicrobial" = alpha("#DE1F84", 0.7), "Not Antimicrobial" = alpha("#C5C5C5", 0.7))) +
theme_void() +
#coord_fixed(ratio = 0.4) +
theme(legend.position = "bottom",
axis.ticks.x = element_blank(),
axis.text.x = element_blank(),
axis.ticks.y = element_blank(),
axis.text.y = element_blank(),
text=element_text(size=10),
panel.background = element_rect(fill = "transparent", color=NA),
plot.background = element_rect(fill = "transparent", colour = NA)) +
labs(x="UMAP 1",
y="UMAP 2",
color="Predicted Activity")
m.umap.over10.annot
# Add information of broad-spectrum activity
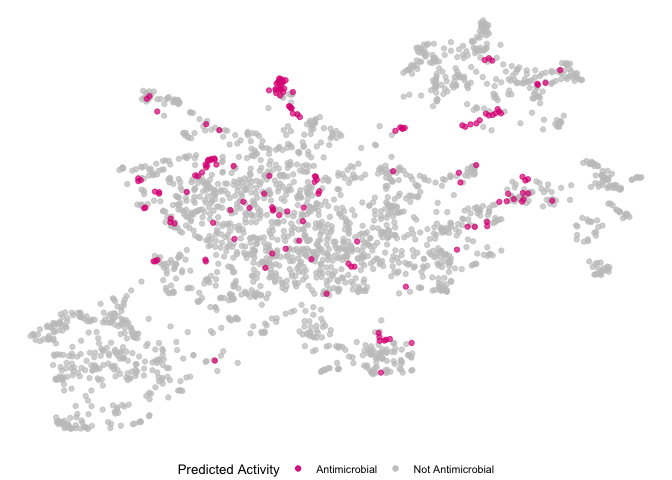
ecfp4.umap <- ecfp4.umap %>%
mutate(pred_activity = if_else(catalog_number %in% ecfp4_predictions.over10$`Catalog Number`, "Antimicrobial", "Not Antimicrobial"))
# Plot
e.umap.over10 <- ggplot(ecfp4.umap, aes(x=V1, y=V2)) +
geom_point(data=subset(ecfp4.umap, pred_activity == "Not Antimicrobial"), aes(color="Not Antimicrobial"), size=1.5) +
geom_point(data=subset(ecfp4.umap, pred_activity == "Antimicrobial"), aes(color="Antimicrobial"), size=1.5) +
scale_color_manual(values=c("Antimicrobial" = alpha("#DE1F84", 0.7), "Not Antimicrobial" = alpha("#C5C5C5", 0.7))) +
theme_void() +
#coord_fixed(ratio = 0.4) +
theme(legend.position = "bottom",
axis.ticks.x = element_blank(),
axis.text.x = element_blank(),
axis.ticks.y = element_blank(),
axis.text.y = element_blank(),
text=element_text(size=10),
panel.background = element_rect(fill = "transparent", color=NA),
plot.background = element_rect(fill = "transparent", colour = NA)) +
labs(x="UMAP 1",
y="UMAP 2",
color="Predicted Activity")
e.umap.over10
chems_interst <- mole_predictions.over10 %>%
filter(ProductName %in% c("Ospemifene", "Shionone", "Bekanamycin", "Doxifluridine", "Ellagic acid")) %>%
select(`Catalog Number`, ProductName) %>%
rename("catalog_number" = "Catalog Number")
chems_interest_umap <- chems_interst %>%
left_join(ecfp4.umap, by="catalog_number")
e.umap.over10.annot <- ggplot(ecfp4.umap, aes(x=V1, y=V2)) +
geom_point(data=subset(ecfp4.umap, pred_activity == "Not Antimicrobial"), aes(color="Not Antimicrobial"), size=1.5) +
geom_point(data=subset(ecfp4.umap, pred_activity == "Antimicrobial"), aes(color="Antimicrobial"), size=1.5) +
geom_text_repel(data = chems_interest_umap, aes(label=ProductName),
max.overlaps = Inf, size=2, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2, fontface="bold") +
scale_color_manual(values=c("Antimicrobial" = alpha("#DE1F84", 0.7), "Not Antimicrobial" = alpha("#C5C5C5", 0.7))) +
theme_void() +
#coord_fixed(ratio = 0.4) +
theme(legend.position = "bottom",
axis.ticks.x = element_blank(),
axis.text.x = element_blank(),
axis.ticks.y = element_blank(),
axis.text.y = element_blank(),
text=element_text(size=10),
panel.background = element_rect(fill = "transparent", color=NA),
plot.background = element_rect(fill = "transparent", colour = NA)) +
labs(x="UMAP 1",
y="UMAP 2",
color="Predicted Activity")
e.umap.over10.annot
ggsave(plot=e.umap.over10.annot,
filename = "../data/05.analyze_mce_predictions/ecfp4_mce_umap.pdf",
width = 21, height = 15, units="cm", dpi=300)
Ranking molecule predictions.
Here we will create plots that compare the antimicrobial activity and the number of predicted inhibited strains.
# Format labels
scatter.nkill.apscore <- mole_predictions %>%
mutate(antibiotic = if_else(antibiotic == "abx", "Antibiotic", "Non-Antibiotic")) %>%
# Create plots
ggplot(aes(x=nk_total, y=apscore_total, color=antibiotic)) +
geom_point(alpha=0.5, size=1) +
scale_color_manual(breaks = c("Antibiotic", "Non-Antibiotic"),
values=c("red", "#C5C5C5")) +
geom_vline(xintercept = 10, linetype="longdash") +
theme_light() +
labs(x="Predicted number of inhibited strains",
y = "Antimicrobial Potential",
color="Compound Class") +
theme(legend.position = c(0.7, 0.3),
text=element_text(size=10),
legend.text = element_text(size=8),
legend.title = element_text(size=8))
score.vs.nkill.marginal <- ggExtra::ggMarginal(scatter.nkill.apscore, type="boxplot", groupColour = TRUE, groupFill = TRUE)
score.vs.nkill.marginal
# Format labels
scatter.nkill.apscore.e <- ecfp4_predictions %>%
mutate(antibiotic = if_else(antibiotic == "abx", "Antibiotic", "Non-Antibiotic")) %>%
# Create plots
ggplot(aes(x=nk_total, y=apscore_total, color=antibiotic)) +
geom_point(alpha=0.5, size=1) +
scale_color_manual(breaks = c("Antibiotic", "Non-Antibiotic"),
values=c("red", "#C5C5C5")) +
geom_vline(xintercept = 10, linetype="longdash") +
theme_light() +
labs(x="Predicted number of inhibited strains",
y = "Antimicrobial Potential",
color="Compound Class") +
theme(legend.position = c(0.7, 0.3),
text=element_text(size=10),
legend.text = element_text(size=8),
legend.title = element_text(size=8))
score.vs.nkill.marginal.e <- ggExtra::ggMarginal(scatter.nkill.apscore.e, type="boxplot", groupColour = TRUE, groupFill = TRUE)
score.vs.nkill.marginal.e
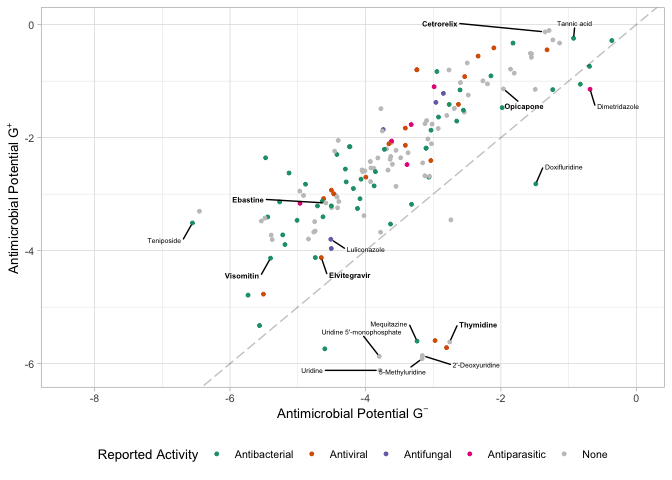
Comparing Gram stains.
Here we compare the antimicrobial potential scores for gram negatives and gram positives.
# Selected chemicals
selected_chems <- c("Visomitin",
"Ebastine",
"Opicapone",
"Cetrorelix (Acetate)",
'Thymidine',
"Elvitegravir")
# Uridine derivatives
uridin.deriv <- c("Uridine",
"Uridine 5'-monophosphate",
"5-Methyluridine",
"2'-Deoxyuridine",
"Doxifluridine")
# Other interests
# Other interests
other.interest <- c("Tannic acid",
"Teniposide",
"Mequitazine",
"Dimetridazole",
"Azelnipidine",
"Luliconazole",
"Bergenin")
# Format Product Names
selected.chemicals.data <- mole_predictions.over10 %>%
mutate(ProductName = if_else(ProductName %in% selected_chems, ProductName, ""),
ProductName = if_else(ProductName == "Cetrorelix (Acetate)", "Cetrorelix", ProductName))
uridine.deriv.data <- mole_predictions.over10 %>%
mutate(ProductName = if_else(ProductName %in% uridin.deriv, ProductName, ""))
other.interest.data <- mole_predictions.over10 %>%
mutate(ProductName = if_else(ProductName %in% other.interest, ProductName, ""))
# Plot
predBS.mole <- mole_predictions.over10 %>%
# Only non-antibiotics
filter(antibiotic == "not_abx") %>%
mutate(`Reported Activity` = if_else(`Reported Activity` %in% c("Antiplasmodium", "Insecticide"), "Antiparasitic", `Reported Activity`)) %>%
ggplot(aes(x=apscore_gnegative, y=apscore_gpositive, color=`Reported Activity`)) +
# Basic aes
geom_point(size=1) +
geom_abline(linetype="longdash", alpha=0.25) +
# Add names
geom_text_repel(data = selected.chemicals.data, aes(x=apscore_gnegative, y=apscore_gpositive, label=ProductName),
max.overlaps = Inf, size=2, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2, fontface="bold",
nudge_x = if_else(selected.chemicals.data$ProductName %in% c("Cetrorelix", "Ebastine"), -1, 0)) +
geom_text_repel(data = uridine.deriv.data, aes(x=apscore_gnegative, y=apscore_gpositive, label=ProductName),
max.overlaps = Inf, size=1.8, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2,
nudge_x = if_else(uridine.deriv.data$ProductName %in% c("Uridine"), -1, 0)) +
geom_text_repel(data = other.interest.data, aes(x=apscore_gnegative, y=apscore_gpositive, label=ProductName),
max.overlaps = Inf, size=1.8, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2) +
# Color by reported activity
scale_color_manual(breaks = c("Antibacterial", "Antiviral", "Antifungal", "Antiparasitic", "None"),
values = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a", "#C5C5C5")) +
theme_light() +
theme(legend.position = "bottom",
text = element_text(size=10)) +
labs(x = latex2exp::TeX("Antimicrobial Potential $G^-$"),
y = latex2exp::TeX("Antimicrobial Potential $G^+$"))
predBS.mole
# Selected chemicals
selected_chems <- c("Visomitin",
"Ebastine",
"Opicapone",
"Cetrorelix (Acetate)",
'Thymidine',
"Elvitegravir")
# Uridine derivatives
uridin.deriv <- c("Uridine",
"Uridine 5'-monophosphate",
"5-Methyluridine",
"2'-Deoxyuridine",
"Doxifluridine")
# Other interests
other.interest <- c("Tannic acid",
"Teniposide",
"Dimetridazole",
"Azelnipidine",
"Luliconazole",
"Bergenin")
# Format Product Names
selected.chemicals.data <- ecfp4_predictions.over10 %>%
mutate(ProductName = if_else(ProductName %in% selected_chems, ProductName, ""),
ProductName = if_else(ProductName == "Cetrorelix (Acetate)", "Cetrorelix", ProductName))
uridine.deriv.data <- ecfp4_predictions.over10 %>%
mutate(ProductName = if_else(ProductName %in% uridin.deriv, ProductName, ""))
other.interest.data <- ecfp4_predictions.over10 %>%
mutate(ProductName = if_else(ProductName %in% other.interest, ProductName, ""))
# Plot
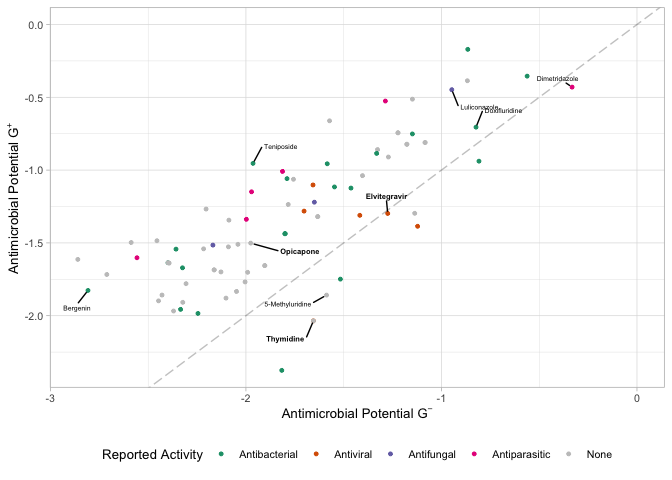
predBS.fps <- ecfp4_predictions.over10 %>%
# Only non-antibiotics
filter(antibiotic == "not_abx") %>%
mutate(`Reported Activity` = if_else(`Reported Activity` %in% c("Antiplasmodium", "Insecticide"), "Antiparasitic", `Reported Activity`)) %>%
ggplot(aes(x=apscore_gnegative, y=apscore_gpositive, color=`Reported Activity`)) +
# Basic aes
geom_point(size=1) +
geom_abline(linetype="longdash", alpha=0.25) +
# Add names
geom_text_repel(data = selected.chemicals.data, aes(x=apscore_gnegative, y=apscore_gpositive, label=ProductName),
max.overlaps = Inf, size=2, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2, fontface="bold",
nudge_x = if_else(selected.chemicals.data$ProductName %in% c("Cetrorelix", "Ebastine"), -1, 0)) +
geom_text_repel(data = uridine.deriv.data, aes(x=apscore_gnegative, y=apscore_gpositive, label=ProductName),
max.overlaps = Inf, size=1.8, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2,
nudge_x = if_else(uridine.deriv.data$ProductName %in% c("Uridine"), -1, 0)) +
geom_text_repel(data = other.interest.data, aes(x=apscore_gnegative, y=apscore_gpositive, label=ProductName),
max.overlaps = Inf, size=1.8, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2) +
# Color by reported activity
scale_color_manual(breaks = c("Antibacterial", "Antiviral", "Antifungal", "Antiparasitic", "None"),
values = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a", "#C5C5C5")) +
theme_light() +
theme(legend.position = "bottom",
text = element_text(size=10)) +
labs(x = latex2exp::TeX("Antimicrobial Potential $G^-$"),
y = latex2exp::TeX("Antimicrobial Potential $G^+$"))
predBS.fps
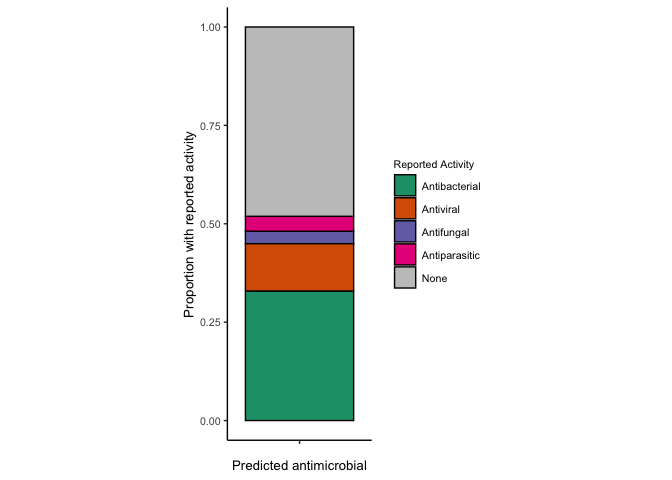
Propotion with reported activity
# Get the activity counts
proportion.known.activity <- mole_predictions.over10 %>%
# Only consider non-antibiotics
filter(antibiotic == "not_abx") %>%
# Consolidate antiparasitic category
mutate(`Reported Activity` = if_else(`Reported Activity` %in% c("Antiplasmodium", "Insecticide"), "Antiparasitic", `Reported Activity`)) %>%
count(`Reported Activity`) %>%
mutate(`Reported Activity` = factor(`Reported Activity`, levels=c("None", "Antiparasitic", "Antifungal", "Antiviral","Antibacterial")),
x=" ")
# Gather
reported.g <- ggplot(proportion.known.activity, aes(x=n, y=x, fill=`Reported Activity`)) +
geom_bar(position="fill", stat="identity", color="black") +
scale_fill_manual(breaks = c("Antibacterial", "Antiviral", "Antifungal", "Antiparasitic", "None"),
values = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a", "#C5C5C5")) +
theme_classic() +
coord_flip() +
theme(text=element_text(size=10),
legend.position = "right",
aspect.ratio = 3,
legend.text = element_text(size = 8),
legend.title = element_text(size=8)) +
labs(x="Proportion with reported activity",
y="Predicted antimicrobial",
fill = "Reported Activity")
reported.g
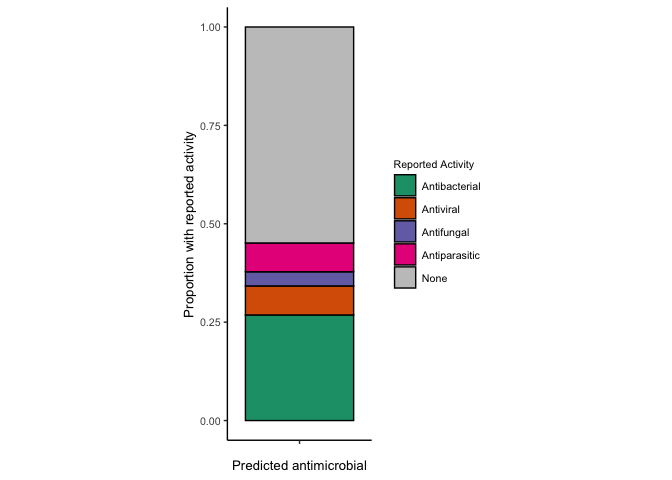
# Get the activity counts
proportion.known.activity.e <- ecfp4_predictions.over10 %>%
# Only consider non-antibiotics
filter(antibiotic == "not_abx") %>%
# Consolidate antiparasitic category
mutate(`Reported Activity` = if_else(`Reported Activity` %in% c("Antiplasmodium", "Insecticide"), "Antiparasitic", `Reported Activity`)) %>%
count(`Reported Activity`) %>%
mutate(`Reported Activity` = factor(`Reported Activity`, levels=c("None", "Antiparasitic", "Antifungal", "Antiviral","Antibacterial")),
x=" ")
# Gather
reported.g.e <- ggplot(proportion.known.activity.e, aes(x=n, y=x, fill=`Reported Activity`)) +
geom_bar(position="fill", stat="identity", color="black") +
scale_fill_manual(breaks = c("Antibacterial", "Antiviral", "Antifungal", "Antiparasitic", "None"),
values = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a", "#C5C5C5")) +
theme_classic() +
coord_flip() +
theme(text=element_text(size=10),
legend.position = "right",
aspect.ratio = 3,
legend.text = element_text(size = 8),
legend.title = element_text(size=8)) +
labs(x="Proportion with reported activity",
y="Predicted antimicrobial",
fill = "Reported Activity")
reported.g.e
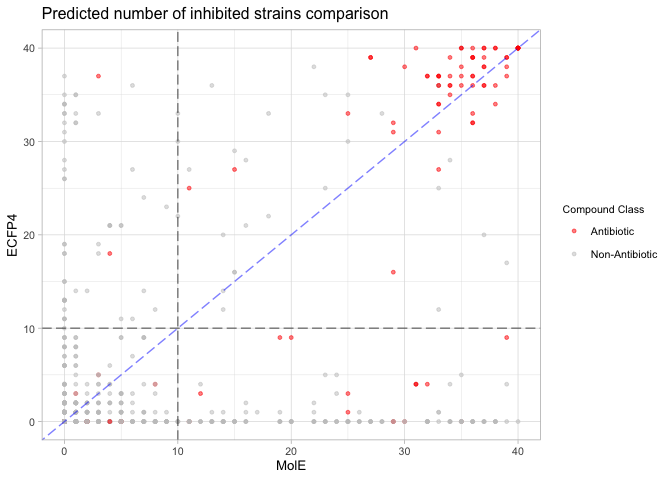
Comparing MolE and ECFP4 predictions.
mole_complete_comp <- mole_predictions %>%
select(`Catalog Number`, nk_total, apscore_total, antibiotic, ProductName) %>%
rename("mole_nk_total" = "nk_total",
"mole_apscore_total" = "apscore_total")
ecfp4_complete_comp <- ecfp4_predictions %>%
select(`Catalog Number`, nk_total, apscore_total) %>%
rename("ecfp4_nk_total" = "nk_total",
"ecfp4_apscore_total" = "apscore_total")
complete_pred_comparison <- ecfp4_complete_comp %>%
left_join(mole_complete_comp, by='Catalog Number') %>%
mutate(antibiotic = if_else(antibiotic == "abx", "Antibiotic", "Non-Antibiotic"))
ggplot(complete_pred_comparison, aes(x=mole_nk_total, y=ecfp4_nk_total, color=antibiotic)) +
geom_point(alpha=0.5, size=1) +
geom_abline(linetype="longdash", alpha=0.5, color="blue") +
geom_vline(xintercept = 10,linetype="longdash", alpha=0.5, color="black") +
geom_hline(yintercept = 10,linetype="longdash", alpha=0.5, color="black") +
scale_color_manual(breaks = c("Antibiotic", "Non-Antibiotic"),
values=c("red", "#C5C5C5")) +
theme_light() +
labs(title="Predicted number of inhibited strains comparison",
x="MolE",
y="ECFP4",
color="Compound Class") +
theme(text=element_text(size=10),
legend.text = element_text(size=8),
legend.title = element_text(size=8))
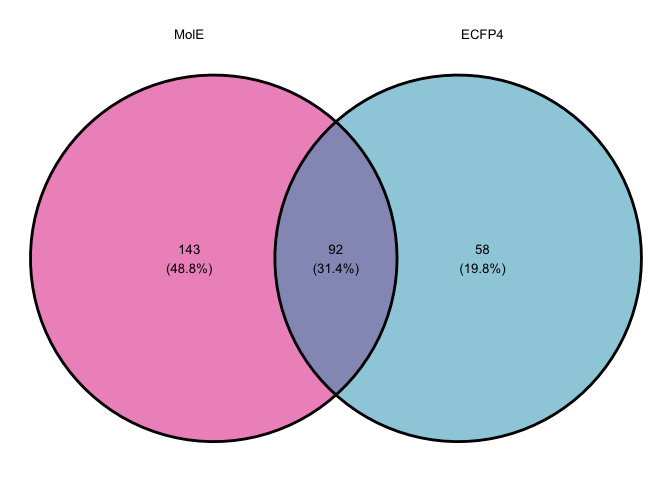
Comparing predicted broad spectrum compounds.
library(ggvenn)
## Loading required package: grid
mole_predictions.over10
## # A tibble: 235 × 13
## `Catalog Number` nk_total nk_gnegative nk_gpositive apscore_total
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 HY-N0797 14 3 11 -3.88
## 2 HY-N0835 13 3 10 -4.16
## 3 HY-P0288 13 2 11 -3.77
## 4 HY-P1564A 25 9 16 -2.95
## 5 HY-D0186 14 11 3 -4.64
## 6 HY-N2177 11 4 7 -4.27
## 7 HY-W009444 14 12 2 -4.67
## 8 HY-16938 16 5 11 -3.29
## 9 HY-13534A 37 15 22 -0.637
## 10 HY-B1916 38 17 21 -0.342
## # ℹ 225 more rows
## # ℹ 8 more variables: apscore_gnegative <dbl>, apscore_gpositive <dbl>,
## # antibiotic <chr>, ProductName <chr>, atc_description <chr>,
## # `Biological Activity` <chr>, `Reported Activity` <chr>, Reference <chr>
fsize <- 3.5
# Complete comparison of broad spectrum antimicrobials
broad.list <- list("MolE" = mole_predictions.over10$`Catalog Number`,
"ECFP4" = ecfp4_predictions.over10$`Catalog Number`)
venn.abx.comparison <- ggvenn(broad.list,
fill_color = c("#DE1F84", "#1F9DBB"),
set_name_size=fsize,
text_size=fsize)
venn.abx.comparison
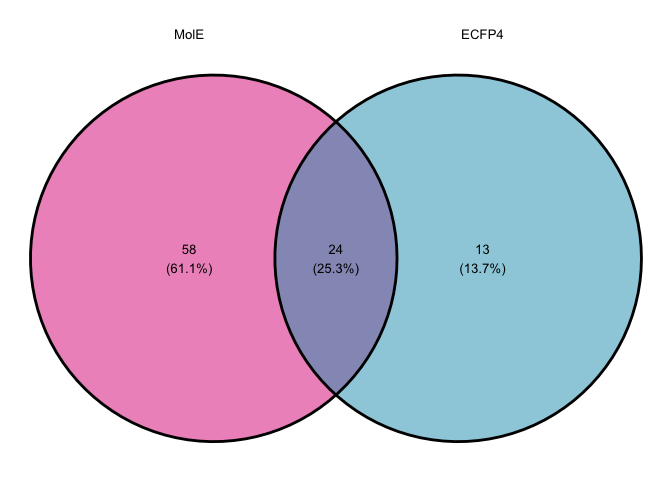
# Comparison of broad spectrum antimicrobials no antibiotics
broad.list <- list("MolE" = mole_predictions.over10 %>% filter(antibiotic == "not_abx",
`Reported Activity` != "None") %>% select(`Catalog Number`) %>% unlist(),
"ECFP4" = ecfp4_predictions.over10 %>% filter(antibiotic == "not_abx",
`Reported Activity` != "None") %>% select(`Catalog Number`) %>% unlist())
venn.nonabx.comparison <- ggvenn(broad.list,
fill_color = c("#DE1F84", "#1F9DBB"),
set_name_size=fsize,
text_size=fsize)
venn.nonabx.comparison
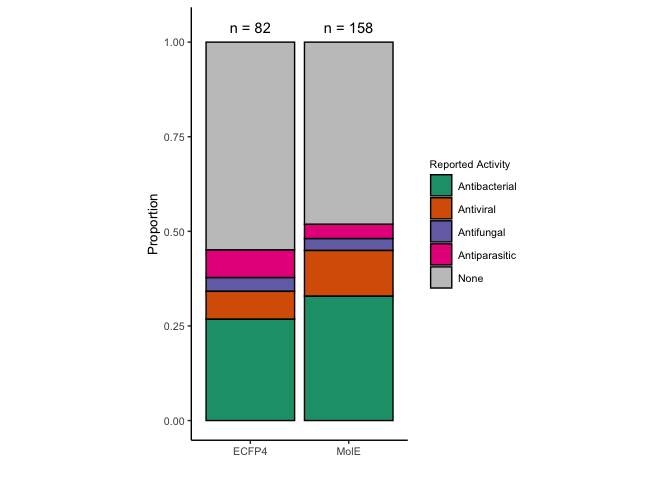
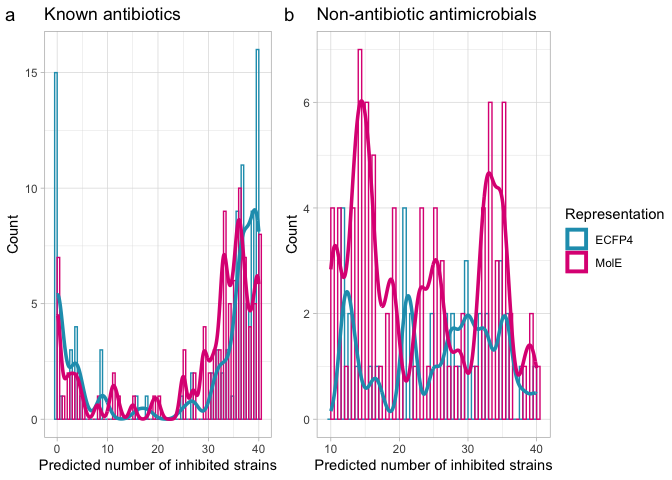
Now compare results from literature search.
proportion.known.activity <- proportion.known.activity %>%
mutate(representation = "MolE")
proportion.known.activity.e <- proportion.known.activity.e %>%
mutate(representation = "ECFP4")
proportion.comparison <- bind_rows(proportion.known.activity,
proportion.known.activity.e)
# Gather totals for n
total_count = proportion.comparison %>%
group_by(representation) %>%
summarise(total = sum(n)) %>%
mutate(tx_sum = paste("n =", total))
reported.bar.comparison <- ggplot(proportion.comparison, aes(x=n, y=representation)) +
annotate("text", y=c("ECFP4", "MolE"), x=1.04, label=total_count$tx_sum,) +
geom_bar(position="fill", stat="identity", color="black", aes(fill=`Reported Activity`)) +
scale_fill_manual(breaks = c("Antibacterial", "Antiviral", "Antifungal", "Antiparasitic", "None"),
values = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a", "#C5C5C5")) +
theme_classic() +
coord_flip() +
theme(text=element_text(size=10),
legend.position = "right",
aspect.ratio = 2,
legend.text = element_text(size = 8),
legend.title = element_text(size=8)) +
labs(y="",
x="Proportion",
fill = "Reported Activity")
reported.bar.comparison
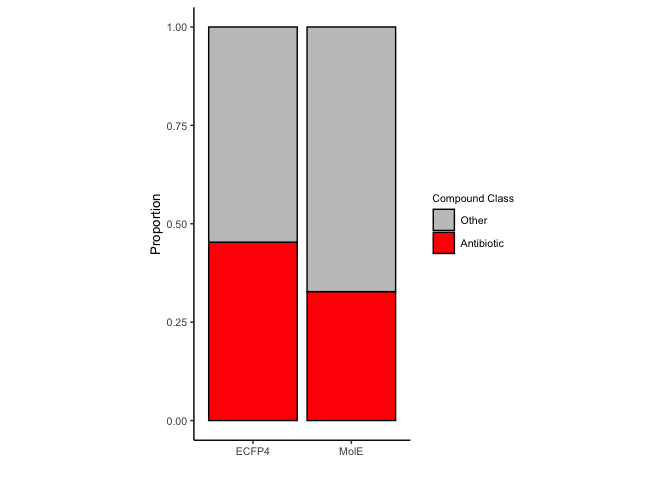
mole.o10.abx <- mole_predictions.over10 %>%
count(antibiotic) %>%
mutate(representation = "MolE")
ecfp4.o10.abx <- ecfp4_predictions.over10 %>%
count(antibiotic) %>%
mutate(representation = "ECFP4")
over10.abx <- bind_rows(mole.o10.abx, ecfp4.o10.abx) %>%
mutate(antibiotic = if_else(antibiotic == "abx", "Antibiotic", "Other"),
antibiotic = factor(antibiotic, levels=c("Other", "Antibiotic")))
abx.bar.comparison <- ggplot(over10.abx, aes(x=n, y=representation, fill=antibiotic)) +
geom_bar(position="fill", stat="identity", color="black") +
scale_fill_manual(breaks = c("Other", "Antibiotic"),
values = c("#C5C5C5", "red")) +
theme_classic() +
coord_flip() +
theme(text=element_text(size=10),
legend.position = "right",
aspect.ratio = 2,
legend.text = element_text(size = 8),
legend.title = element_text(size=8)) +
labs(y="",
x="Proportion",
fill = "Compound Class")
abx.bar.comparison
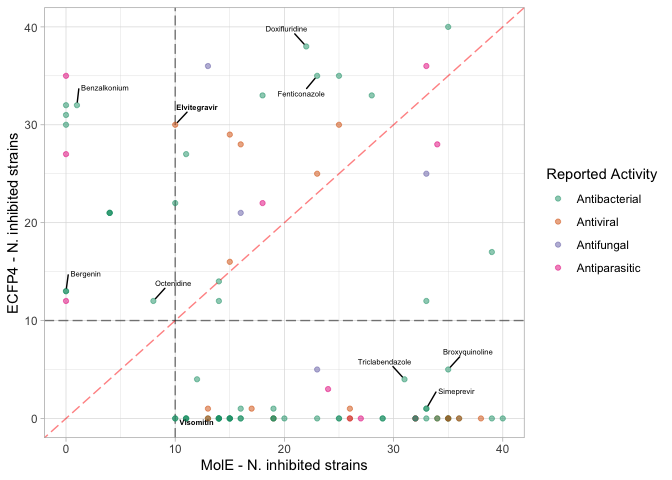
Comparing predicted broad spectrum.
mole.litsearch <- mole_predictions.over10 %>%
filter(antibiotic == "not_abx") %>%
select(`Catalog Number`, `Reported Activity`, ProductName) %>%
# Consolidate antiparasitic category
mutate(`Reported Activity` = if_else(`Reported Activity` %in% c("Antiplasmodium", "Insecticide"), "Antiparasitic", `Reported Activity`))
ecfp4.litsearch <- ecfp4_predictions.over10 %>%
filter(antibiotic == "not_abx") %>%
select(`Catalog Number`, `Reported Activity`, ProductName) %>%
# Consolidate antiparasitic category
mutate(`Reported Activity` = if_else(`Reported Activity` %in% c("Antiplasmodium", "Insecticide"), "Antiparasitic", `Reported Activity`))
combined.litsearch <- bind_rows(mole.litsearch, ecfp4.litsearch) %>%
distinct()
mole_o10 <- mole_predictions %>%
filter(`Catalog Number` %in% combined.litsearch$`Catalog Number`) %>%
select(`Catalog Number`, nk_total, apscore_total) %>%
rename("mole_nk_total" = "nk_total",
"mole_apscore_total" = "apscore_total")
ecfp4_o10 <- ecfp4_predictions %>%
filter(`Catalog Number` %in% combined.litsearch$`Catalog Number`) %>%
select(`Catalog Number`, nk_total, apscore_total) %>%
rename("ecfp4_nk_total" = "nk_total",
"ecfp4_apscore_total" = "apscore_total")
broadS.comp <- combined.litsearch %>%
left_join(mole_o10, by="Catalog Number") %>%
left_join(ecfp4_o10, by="Catalog Number") %>%
filter(`Reported Activity` != "None")
# Selected chemicals
selected_chems <- c("Visomitin",
"Ebastine",
"Opicapone",
"Cetrorelix (Acetate)",
'Thymidine',
"Elvitegravir")
# Other interests
other.interest <- c("Bergenin",
"Fenticonazole (Nitrate)",
"Doxifluridine",
"Broxyquinoline",
"Triclabendazole",
"Octenidine (dihydrochloride)",
"Benzalkonium (chloride)",
"Simeprevir")
# Format Product Names
selected.chemicals.data <- broadS.comp %>%
mutate(ProductName = if_else(ProductName %in% selected_chems, ProductName, ""),
ProductName = if_else(ProductName == "Cetrorelix (Acetate)", "Cetrorelix", ProductName))
other.interest.data <- broadS.comp %>%
mutate(ProductName = if_else(ProductName %in% other.interest, ProductName, ""),
ProductName = if_else(ProductName == "Fenticonazole (Nitrate)", "Fenticonazole", ProductName),
ProductName = if_else(ProductName == "Octenidine (dihydrochloride)", "Octenidine", ProductName),
ProductName = if_else(ProductName == "Benzalkonium (chloride)", "Benzalkonium", ProductName))
nk.comparison <- ggplot(broadS.comp, aes(x=mole_nk_total, y=ecfp4_nk_total, color=`Reported Activity`)) +
geom_vline(xintercept = 10, linetype="longdash", color="black", alpha=0.5) +
geom_hline(yintercept=10, linetype="longdash", color="black", alpha=0.5) +
geom_point(alpha=0.5) +
geom_abline(linetype="longdash", color="red", alpha=0.5) +
# Add names
geom_text_repel(data = selected.chemicals.data, aes(x=mole_nk_total, y=ecfp4_nk_total, label=ProductName),
max.overlaps = Inf, size=2, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2, fontface="bold",
nudge_x = if_else(selected.chemicals.data$ProductName %in% c("Elvitegravir"), 1, 0)) +
geom_text_repel(data = other.interest.data, aes(x=mole_nk_total, y=ecfp4_nk_total, label=ProductName),
max.overlaps = Inf, size=2, min.segment.length = 0, color="black", box.padding = 0.7, point.padding = 0.2,
nudge_y = if_else(other.interest.data$ProductName %in% c("Benzalkonium"), 1, 0)) +
scale_color_manual(breaks = c("Antibacterial", "Antiviral", "Antifungal", "Antiparasitic", "None"),
values = c("#1b9e77", "#d95f02", "#7570b3", "#e7298a", "#C5C5C5")) +
theme_light() +
labs(x="MolE - N. inhibited strains",
y="ECFP4 - N. inhibited strains")
nk.comparison
Figure panels.
library(ggpubr)
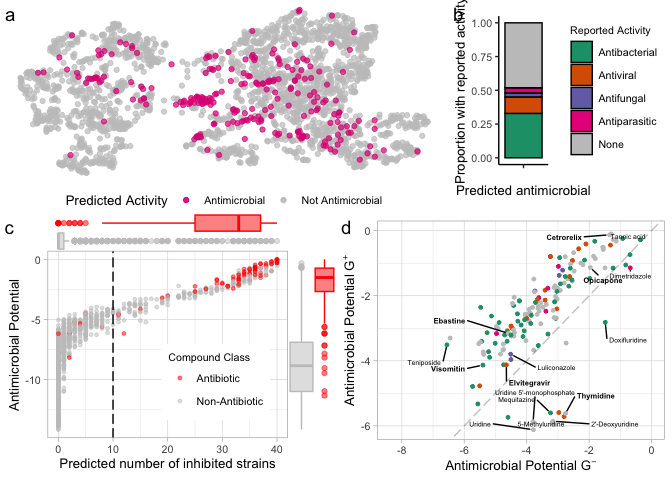
top_row <- ggarrange(e.umap.over10 + theme(aspect.ratio = 0.4), reported.g.e, labels = c("a", "b"),
widths = c(2, 1),font.label = list(face="plain"))
bottom_row <- ggarrange(score.vs.nkill.marginal.e, predBS.fps + theme(legend.position = "none"),
labels = c("c", "d"), font.label = list(face="plain"))
pred.fps.panel <- ggarrange(top_row, bottom_row, ncol = 1, nrow = 2, heights = c(0.8, 1))
pred.fps.panel
ggsave(filename = "../data/05.analyze_mce_predictions/ecfp4_mce_overview.pdf",
plot=pred.fps.panel,
width = 21, height = 15, units="cm", dpi=300)
ggsave(filename = "../data/05.analyze_mce_predictions/ecfp4_mce_overview.svg",
plot=pred.fps.panel,
width = 21, height = 15, units="cm", dpi=300)
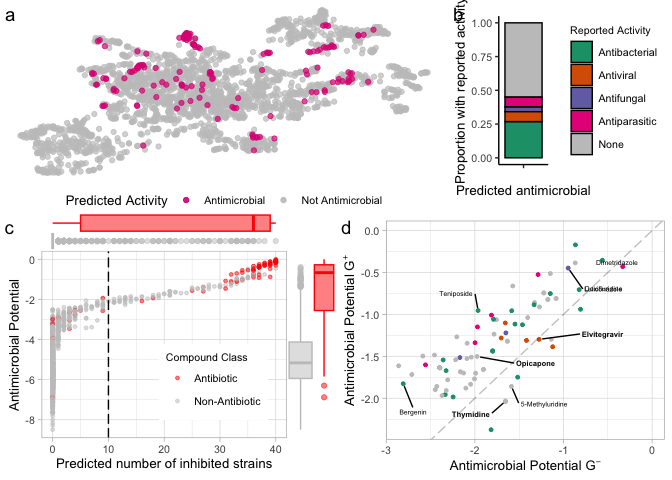
top_row <- ggarrange(m.umap.over10 + theme(aspect.ratio = 0.4), reported.g, labels = c("a", "b"),
widths = c(2, 1),font.label = list(face="plain"))
bottom_row <- ggarrange(score.vs.nkill.marginal, predBS.mole + theme(legend.position = "none"),
labels = c("c", "d"), font.label = list(face="plain"))
pred.mole.panel <- ggarrange(top_row, bottom_row, ncol = 1, nrow = 2, heights = c(0.8, 1))
pred.mole.panel
ggsave(filename = "../data/05.analyze_mce_predictions/mole_mce_overview.pdf",
plot=pred.mole.panel,
width = 21, height = 15, units="cm", dpi=300)
Combine complete prediction information
ecfp4_predranks <- ecfp4_predictions %>%
mutate(representation = "ECFP4")
mole_predranks <- mole_predictions %>%
mutate(representation = "MolE")
complete_predranks <- bind_rows(ecfp4_predranks, mole_predranks)
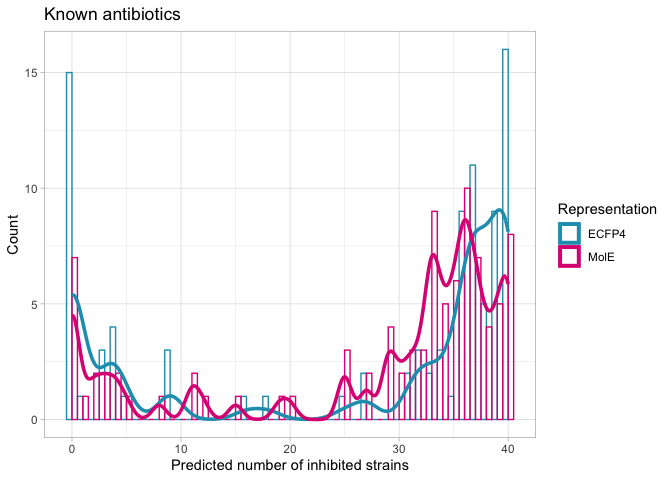
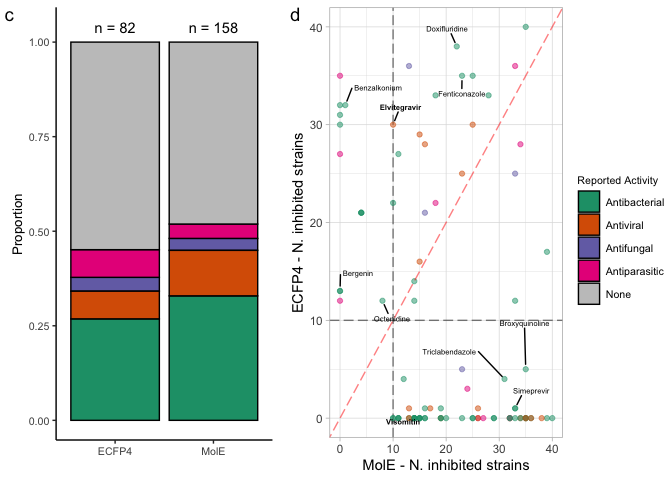
nk.abx.hist <- complete_predranks %>%
filter(antibiotic == "abx") %>%
ggplot(aes(x=nk_total, color=representation, color=representation)) +
geom_histogram(binwidth = 1, aes(y=after_stat(count)), position="dodge", alpha=0.1, fill="white") +
geom_density(alpha=0.2, aes(y=after_stat(count)), size=1.25, adjust=1/5) +
scale_color_manual(values=c("#1F9DBB", "#DE1F84")) +
theme_light() +
labs(x="Predicted number of inhibited strains",
y="Count",
color="Representation",
title = "Known antibiotics")
## Warning: Duplicated aesthetics after name standardisation: colour
## Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
## ℹ Please use `linewidth` instead.
## This warning is displayed once every 8 hours.
## Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
## generated.
nk.abx.hist
mole.o10 <- mole_predictions.over10 %>%
filter(antibiotic == "not_abx") %>%
filter(`Reported Activity` != "None") %>%
select(`Catalog Number`) %>%
unlist()
ecfp4.o10 <- ecfp4_predictions.over10 %>%
filter(antibiotic == "not_abx") %>%
filter(`Reported Activity` != "None") %>%
select(`Catalog Number`) %>%
unlist()
ecfp4.comppreds <- complete_predranks %>%
filter(representation == "ECFP4",
`Catalog Number` %in% ecfp4.o10)
mole.comppreds <- complete_predranks %>%
filter(representation == "MolE",
`Catalog Number` %in% mole.o10)
complete_litsearch_comparison <- bind_rows(ecfp4.comppreds, mole.comppreds)
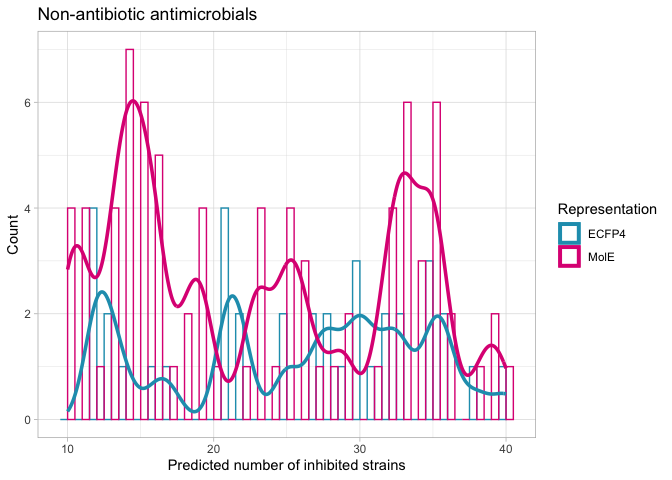
nk.ls.hist <- ggplot(complete_litsearch_comparison, aes(x=nk_total, color=representation)) +
geom_histogram(binwidth = 1, aes(y=after_stat(count)), position="dodge", alpha=0.1, fill="white") +
geom_density(alpha=0.2, aes(y=after_stat(count)), size=1.25, adjust=1/4) +
scale_color_manual(values=c("#1F9DBB", "#DE1F84")) +
#scale_fill_manual(values=c("#1F9DBB", "#DE1F84")) +
theme_light() +
labs(x="Predicted number of inhibited strains",
y="Count",
color="Representation",
title = "Non-antibiotic antimicrobials")
nk.ls.hist
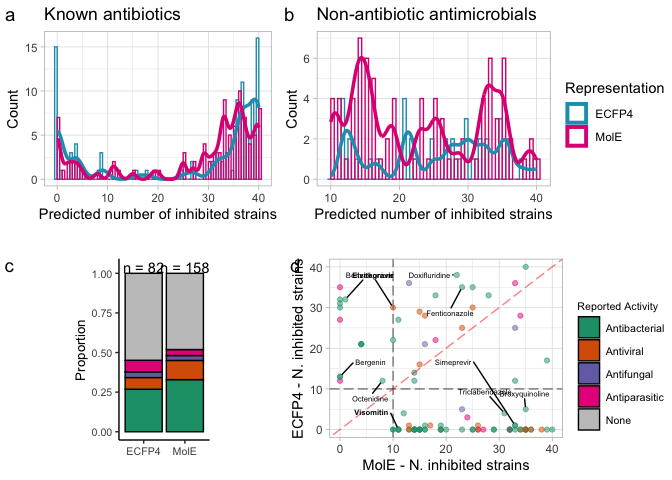
panel.comp.nk <- ggarrange(nk.abx.hist, nk.ls.hist,
nrow = 1, ncol = 2,
labels = c("a", "b"),
font.label = list(face="plain"),
common.legend = TRUE,
legend = "right")
panel.comp.nk
panel.comp.lit <- ggarrange(reported.bar.comparison, nk.comparison,
nrow = 1, ncol = 2,
labels = c("c", "d"),
font.label = list(face="plain"),
common.legend = TRUE,
legend = "right")
panel.comp.lit
complete.comp.panel <- ggarrange(panel.comp.nk,
NULL,
panel.comp.lit,
heights = c(1, 0.1, 1),
nrow = 3, ncol = 1)
ggsave(plot = complete.comp.panel,
filename = "../data/05.analyze_mce_predictions/mce_pred_comparison.pdf",
width = 21, height = 18, units="cm", dpi=300)
ggsave(plot = complete.comp.panel,
filename = "../data/05.analyze_mce_predictions/mce_pred_comparison.png",
width = 21, height = 18, units="cm", dpi=300)
complete.comp.panel
Session Info
sessionInfo()
## R version 4.3.1 (2023-06-16)
## Platform: x86_64-apple-darwin20 (64-bit)
## Running under: macOS Ventura 13.4.1
##
## Matrix products: default
## BLAS: /Library/Frameworks/R.framework/Versions/4.3-x86_64/Resources/lib/libRblas.0.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/4.3-x86_64/Resources/lib/libRlapack.dylib; LAPACK version 3.11.0
##
## locale:
## [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
##
## time zone: Europe/Berlin
## tzcode source: internal
##
## attached base packages:
## [1] grid stats graphics grDevices utils datasets methods
## [8] base
##
## other attached packages:
## [1] ggpubr_0.6.0 ggvenn_0.1.10 uwot_0.1.16 Matrix_1.6-1.1
## [5] readxl_1.4.2 ggrepel_0.9.3 lubridate_1.9.2 forcats_1.0.0
## [9] stringr_1.5.0 dplyr_1.1.2 purrr_1.0.2 readr_2.1.4
## [13] tidyr_1.3.0 tibble_3.2.1 ggplot2_3.4.4 tidyverse_2.0.0
##
## loaded via a namespace (and not attached):
## [1] tidyselect_1.2.0 farver_2.1.1 fastmap_1.1.1 latex2exp_0.9.6
## [5] promises_1.2.0.1 digest_0.6.33 timechange_0.2.0 mime_0.12
## [9] lifecycle_1.0.4 ellipsis_0.3.2 magrittr_2.0.3 compiler_4.3.1
## [13] rlang_1.1.4 tools_4.3.1 utf8_1.2.4 yaml_2.3.7
## [17] knitr_1.43 ggsignif_0.6.4 labeling_0.4.3 bit_4.0.5
## [21] abind_1.4-5 miniUI_0.1.1.1 withr_2.5.2 fansi_1.0.6
## [25] xtable_1.8-4 colorspace_2.1-0 scales_1.3.0 cli_3.6.3
## [29] rmarkdown_2.24 crayon_1.5.2 ragg_1.2.5 generics_0.1.3
## [33] rstudioapi_0.15.0 tzdb_0.4.0 parallel_4.3.1 cellranger_1.1.0
## [37] vctrs_0.6.5 carData_3.0-5 car_3.1-2 hms_1.1.3
## [41] bit64_4.0.5 rstatix_0.7.2 irlba_2.3.5.1 systemfonts_1.0.4
## [45] glue_1.7.0 codetools_0.2-19 cowplot_1.1.1 RcppAnnoy_0.0.21
## [49] stringi_1.7.12 gtable_0.3.4 later_1.3.1 munsell_0.5.0
## [53] pillar_1.9.0 htmltools_0.5.6 ggExtra_0.10.0 R6_2.5.1
## [57] textshaping_0.3.6 vroom_1.6.3 evaluate_0.21 shiny_1.7.4
## [61] lattice_0.21-8 highr_0.10 backports_1.4.1 broom_1.0.5
## [65] httpuv_1.6.11 Rcpp_1.0.13 gridExtra_2.3 svglite_2.1.1
## [69] xfun_0.40 pkgconfig_2.0.3